Abstract
Deep learning in de novo protein design has achieved atomic-level fidelity. However, existing models remain largely non-deliberative: they directly synthesize molecular geometries without explicitly reasoning about which residues or interactions are functionally essential. As a result, design decisions are entangled with continuous sampling dynamics, limiting interpretability, controllability, and systematic reuse of biochemical knowledge. We introduce Proteo-R1, a reasoning-guided protein design framework that explicitly decouples molecular understanding from geometric generation. Proteo-R1 adopts a dual-expert architecture in which a multimodal large language model (MLLM) serves as an understanding expert, analyzing protein sequences, structures, and textual context to identify key functional residues that govern binding and specificity. These residue-level decisions are then passed as hard constraints to a separate diffusion-based generation expert, which performs conditional co-design while respecting fixed interaction anchors. This factorization mirrors how human experts approach molecular engineering: first, reasoning about critical interactions, then optimizing geometry subject to those constraints. By operationalizing reasoning as explicit residue-level commitments rather than latent textual guidance, Proteo-R1 achieves stable, interpretable, and modular integration of LLM reasoning with state-of-the-art geometric generative models.
Method Overview
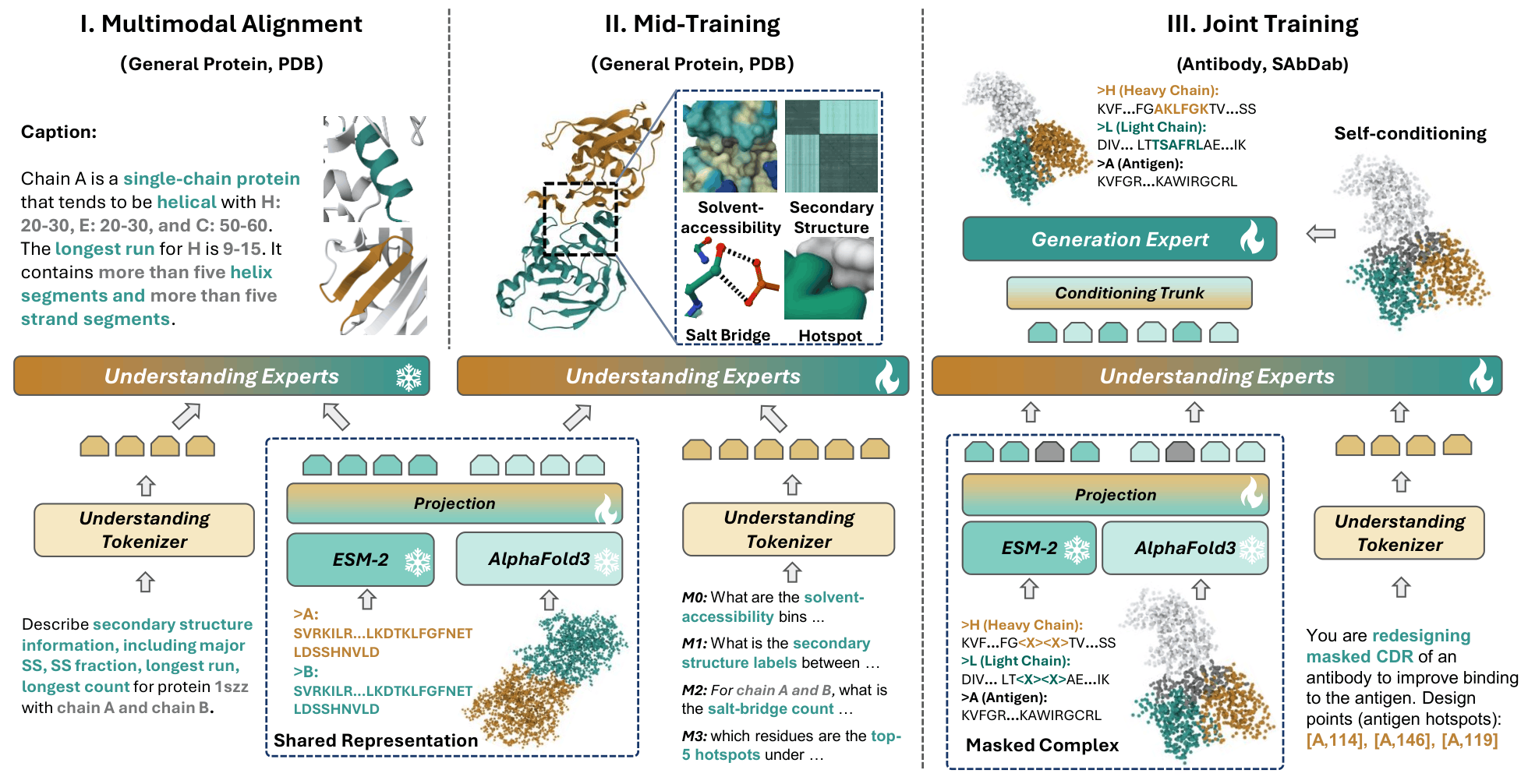
Proteo-R1 is a dual-expert framework that couples a multimodal reasoning expert with a diffusion-based generation expert. The reasoner identifies residue-level interaction anchors from sequence, structure, and instruction context, and the generator performs conditional co-design under these explicit biochemical constraints.
Key Results
- Reasoning and generation are explicitly decoupled via residue-level anchors.
- Reasoning-guided CDR co-design improves realism, controllability, and interpretability.
- The architecture is modular and can integrate with modern geometric generators.
Main Tables from Paper
Geometry-Centric Evaluation of Simultaneous Multi-CDR Redesign
| Method | H1 | H2 | H3 | L1 | L2 | L3 | Loop-RMSD | IMP | Clash_in | Clash_out | JSD_bb |
|---|---|---|---|---|---|---|---|---|---|---|---|
| DiffAb | 1.52 | 1.44 | 4.29 | 1.43 | 1.21 | 1.80 | 5.03 | 53.35 | -- | -- | -- |
| dyMEAN | 1.65 | 1.47 | 6.15 | 1.58 | 1.23 | 1.59 | 7.84 | 5.60 | -- | -- | -- |
| HTP | 1.56 | 1.45 | 4.32 | 1.55 | 1.20 | 1.73 | 7.18 | 6.09 | -- | -- | -- |
| IgGM | 1.73 | 1.55 | 4.37 | 1.62 | 1.51 | 1.71 | 9.18 | 9.01 | 25.63% | 1.45% | 0.2873 |
| AbX | 1.55 | 1.23 | 4.91 | 0.76 | 0.40 | 1.30 | 5.77 | 52.26 | 1.47% | 0.30% | 0.2497 |
| MFDesign | 1.61 | 1.44 | 3.71 | 1.65 | 1.15 | 1.69 | 4.28 | 59.16 | 0.53% | 0.26% | 0.2734 |
| Proteo-R1 | 1.33 | 1.13 | 3.81 | 1.54 | 0.85 | 1.51 | 4.51 | 56.58 | 0.50% | 0.14% | 0.2661 |
CDR-H3 Design on RAbD
| Model | AAR | lDDT | TMscore | RMSD | DockQ |
|---|---|---|---|---|---|
| RosettaAb* | 32.31% | 0.8272 | 0.9717 | 17.70 | 0.137 |
| DiffAb* | 35.31% | 0.8281 | 0.9695 | 23.24 | 0.158 |
| MEAN* | 37.38% | 0.8252 | 0.9688 | 17.30 | 0.162 |
| GeoAB* | 40.02% | 0.8367 | 0.9695 | 15.43 | 0.187 |
| HERN | 32.65% | --- | --- | 9.15 | 0.294 |
| dyMEAN | 41.84% | 0.8392 | 0.9718 | 8.10 | 0.407 |
| DGENet | 42.67% | 0.8551 | 0.9747 | 7.19 | 0.431 |
| BoltzGen | 39.07% | 0.8372 | 0.9675 | 2.69 | 0.473 |
| Proteo-R1 | 10.75% | 0.9693 | 0.9816 | 2.46 | 0.801 |
Sequence Recovery vs Inverse Folding Consistency
| CDR | AbX (AAR / IF-AAR / Delta) | IgGM (AAR / IF-AAR / Delta) | MFDesign (AAR / IF-AAR / Delta) | Proteo-R1 (AAR / IF-AAR / Delta) |
|---|---|---|---|---|
| H1 | 71.34 / 59.80 / -11.54 | 73.98 / 62.76 / -11.22 | 74.95 / 60.90 / -14.05 | 42.62 / 61.17 / +18.55 |
| H2 | 59.15 / 46.10 / -13.05 | 59.15 / 45.79 / -13.36 | 67.54 / 40.63 / -26.91 | 18.97 / 31.67 / +12.70 |
| H3 | 31.58 / 18.96 / -12.62 | 29.42 / 19.55 / -9.87 | 65.04 / 19.73 / -45.31 | 15.06 / 19.27 / +4.21 |
| L1 | 89.13 / 62.02 / -27.11 | 72.20 / 56.53 / -15.67 | 82.98 / 54.94 / -28.04 | 47.12 / 51.40 / +4.28 |
| L2 | 90.90 / 62.33 / -28.57 | 71.43 / 55.66 / -15.77 | 87.81 / 53.22 / -34.59 | 46.43 / 51.43 / +5.00 |
| L3 | 67.82 / 43.49 / -24.33 | 59.43 / 41.84 / -17.59 | 80.15 / 40.98 / -39.17 | 40.43 / 37.09 / -3.34 |
Compatibility with UniMoMo Backbone (CDR-H3)
| Model | #Gen | AAR | RMSD | IMP | Delta G |
|---|---|---|---|---|---|
| MEAN | 1 | 29.13% | 1.87 | 6.67% | -- |
| dyMEAN | 1 | 31.65% | 8.21 | 11.86% | -- |
| GeoAB-R | 1 | 32.04% | 1.67 | 6.67% | -- |
| UniMoMo (all) | 100 | 52.34% | 1.04 | 65.00% | 8.46 |
| Proteo-R1 (UniMoMo) | 100 | 48.94% | 0.83 | 67.79% | 7.35 |
BibTeX
@inproceedings{proteor1_icml2026,
title={Proteo-R1: Reasoning Foundation Models for De Novo Protein Design},
author={Wu, Fang and Xuan, Weihao and Qi, Heli and Cao, Hanqun and Chang, Heng-Jui and Zhou, Zeqi and Zhao, Haokai and Jian, Ma and Ma, Carl and Cheng, Yu-Chi and Pang, Kuan and Tang, Xiangru and Wang, Zehong and Li, Guanlue and Wang, Hanchen and Ying, Kejun and Lu, Pan and Im, Chiho and Han, Seungju and Xia, Peng and Xu, Tinson and Li, Yinxi and Zhu, Deyao and Heng, Pheng-Ann and Yokoya, Naoto and Sugiyama, Masashi and Li, Li Erran and Leskovec, Jure and Choi, Yejin},
booktitle={International Conference on Machine Learning (ICML)},
year={2026}
}