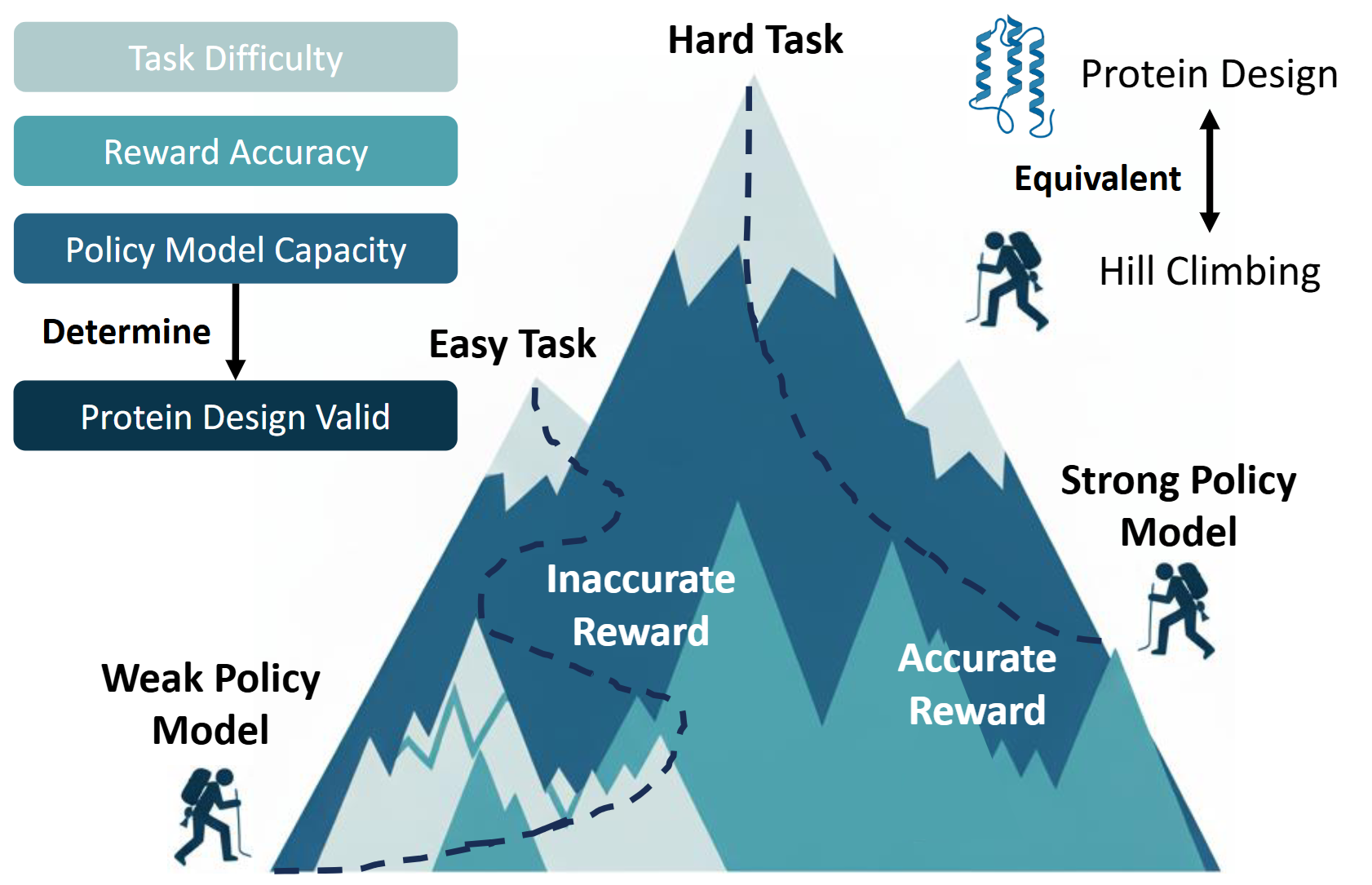
 From Supervision to Exploration: What Does Protein Language Model Learn During Reinforcement Learning?
From Supervision to Exploration: What Does Protein Language Model Learn During Reinforcement Learning?
Hanqun Cao*, Hongrui Zhang*, Junde Xu*, Zhou Zhang*, Lingdong Shen, Minghao Sun, Ge Liu, Jinbo Xu, Wu-Jun Li, Jinren Ni, Cesar de la Fuente-Nunez,
Tianfan Fu, Yejin Choi, Pheng-Ann Heng, Fang Wu†
Under Review
[Paper]
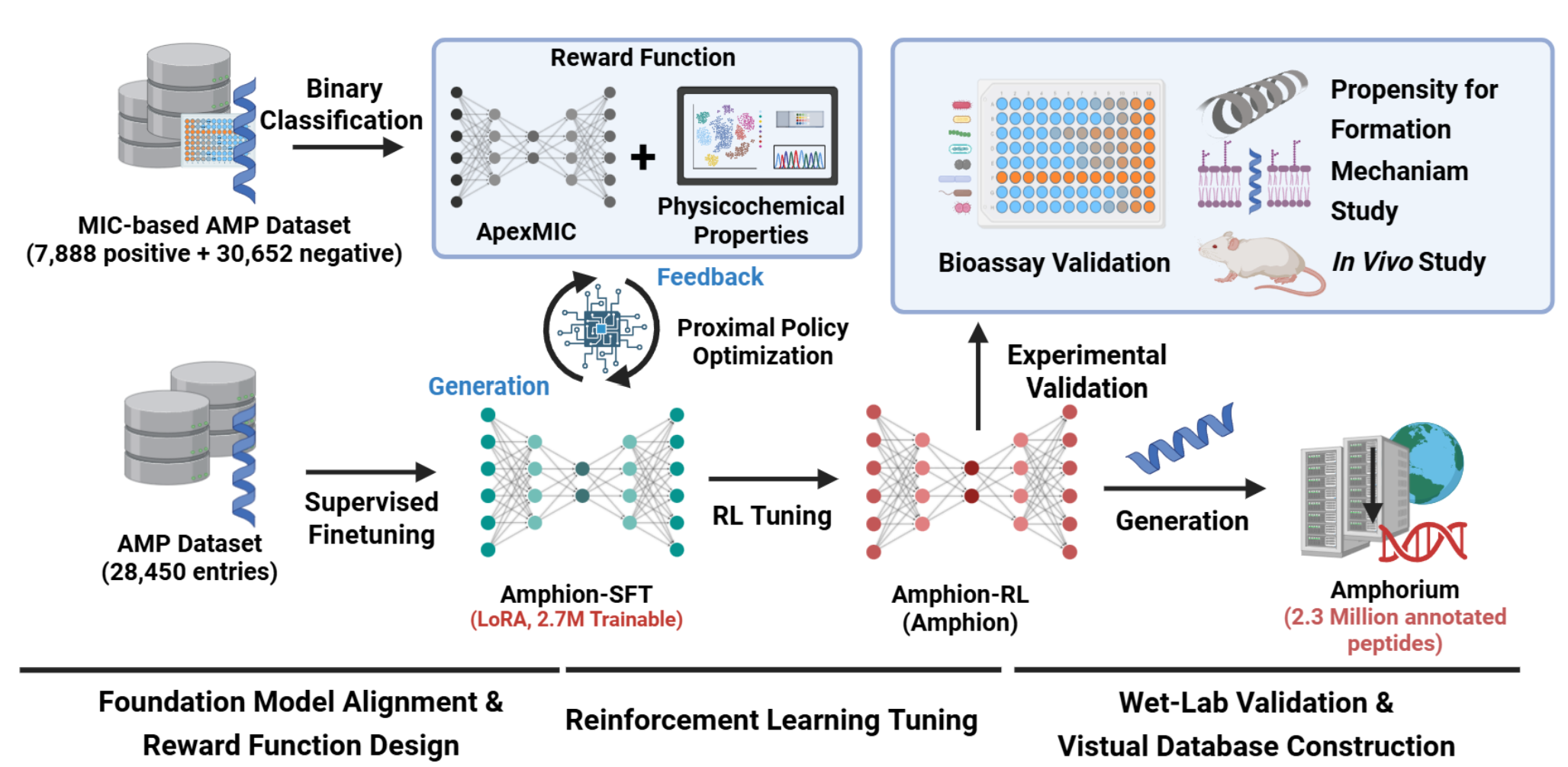 A Deep Reinforcement Learning Platform for Antibiotic Discovery.
A Deep Reinforcement Learning Platform for Antibiotic Discovery.
![GitHub stars]()
Hanqun Cao, Marcelo D. T. Torres, Jingjie Zhang, Zijun Gao, Fang Wu, Chunbin Gu, Jure Leskovec, Yejin Choi,
Cesar de la Fuente-Nunez, Guangyong Chen, Pheng-Ann Heng†
Under Review
[Paper]
[Code]
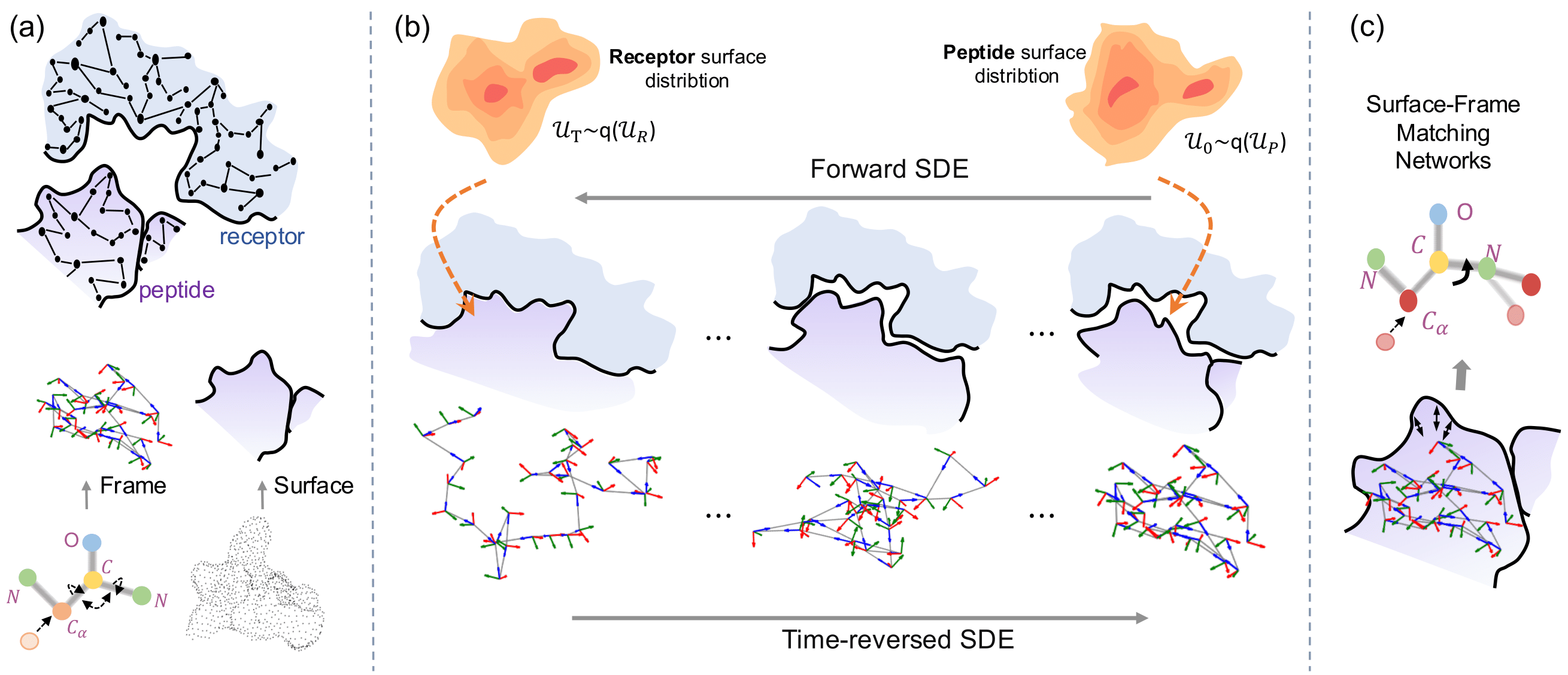 Joint Design of Protein Surface and Structure Using a Diffusion Bridge Model.
Joint Design of Protein Surface and Structure Using a Diffusion Bridge Model.

Guanlue Li, Xufeng Zhao, Fang Wu,† , Soren Laue†
NeurIPS 2025
[Paper]
[Code]
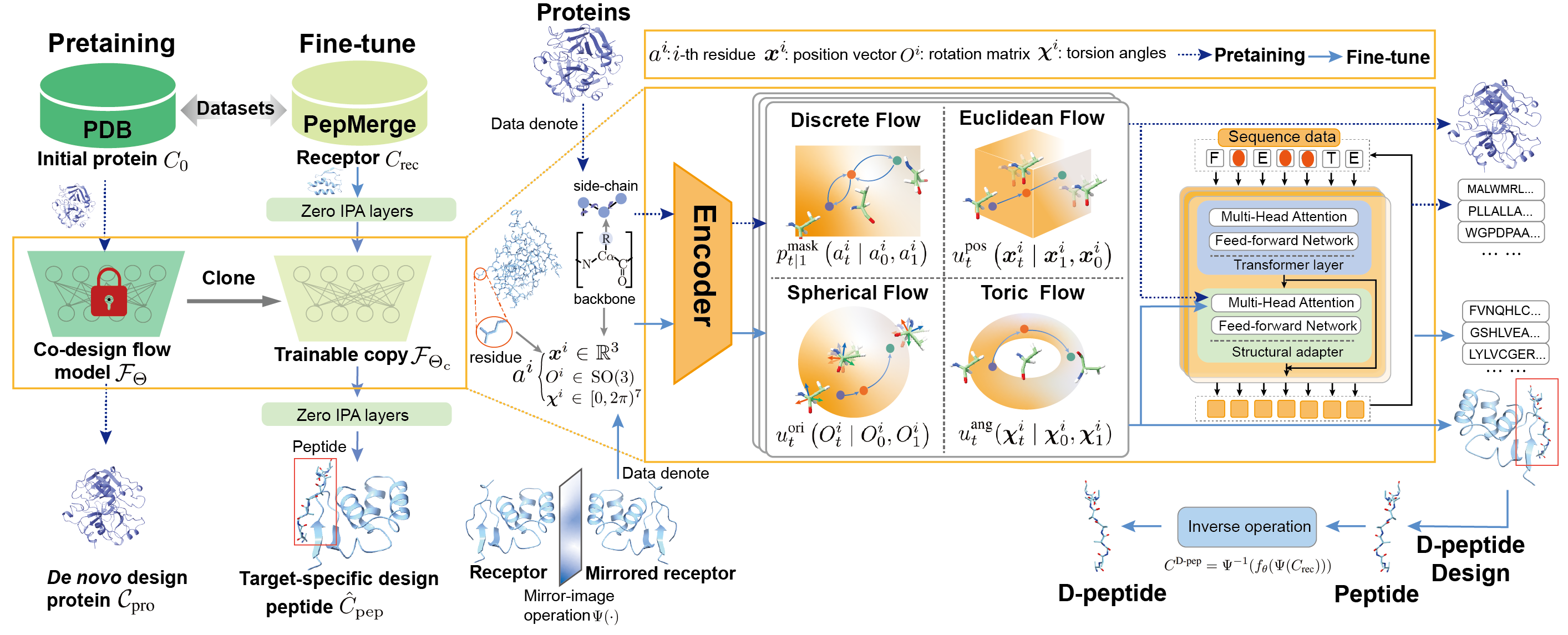 D-Flow: Multi-modality Flow Matching for D-peptide Design.
D-Flow: Multi-modality Flow Matching for D-peptide Design.

Fang Wu*, Tinson Xu*, Shuting Jin*, Xiangru Tang, Zerui Xu, James Zou, Brian Hie†
NeurIPS 2025 AI4D3 Workshop
[Paper]
[Code]
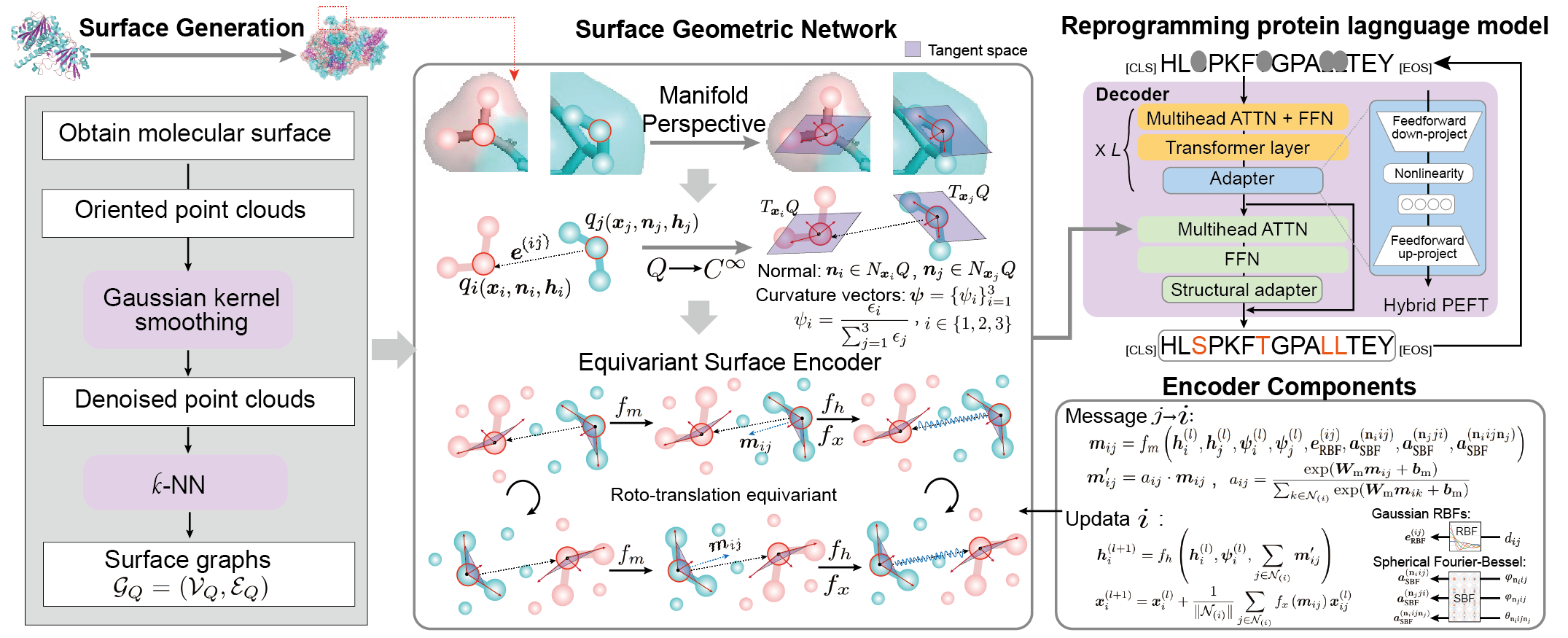 SurfDesign: Effective Protein Design on Molecular Surfaces.
SurfDesign: Effective Protein Design on Molecular Surfaces.
Fang Wu, Shuting Jin, Jianmin Wang, Zerui Xu, xiangxiang Zeng, Jinbo Xu†
Under review
[Paper]
[Code]
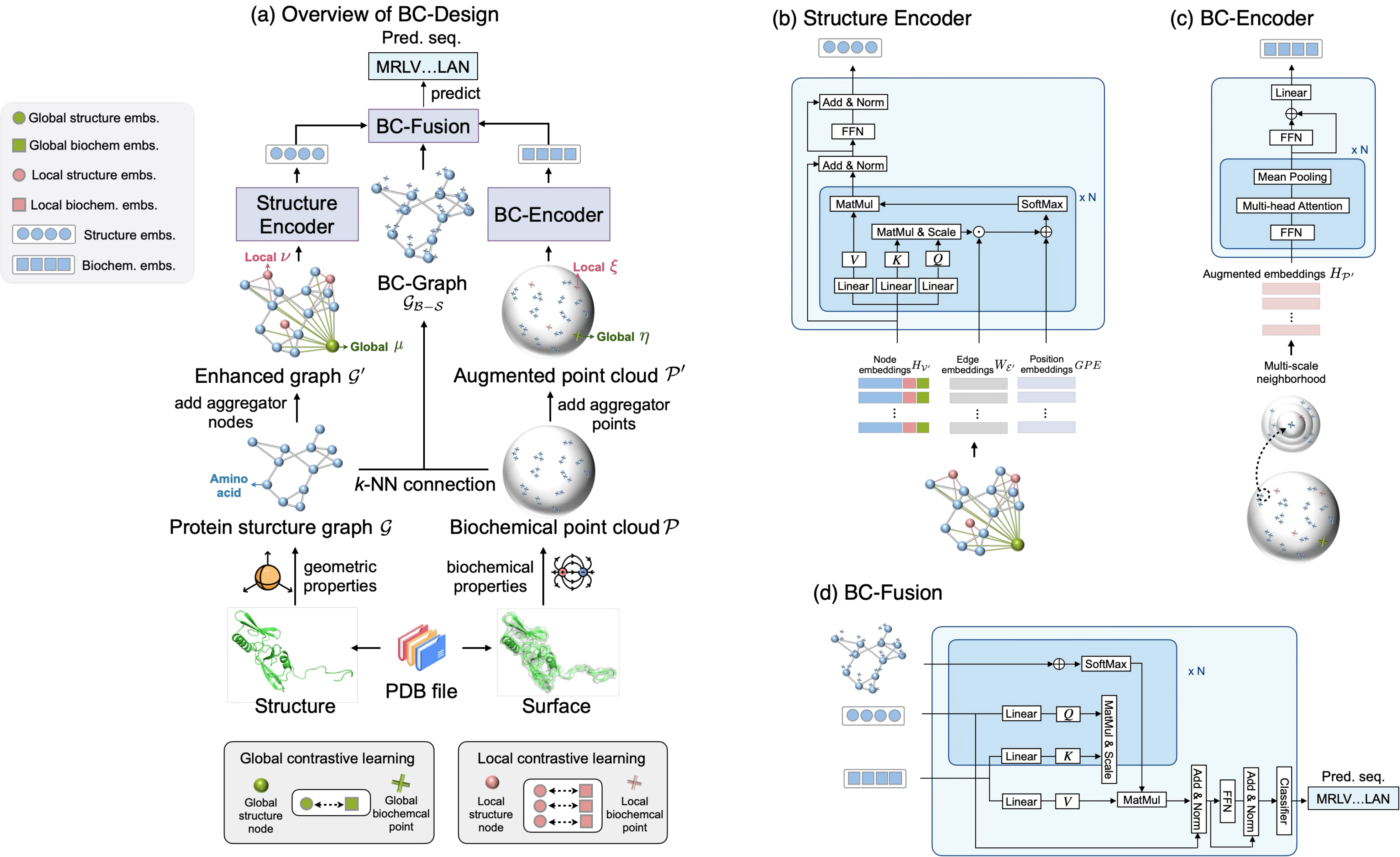 BC-Design: A Biochemistry-Aware Framework for High-Precision Inverse Protein Folding.
BC-Design: A Biochemistry-Aware Framework for High-Precision Inverse Protein Folding.

Xiangru Tang*, Xinwu Ye*, Fang Wu*, Yanjun Shao, Yin Fang, Siming Chen, Dong Xu, Mark Gerstein†
ICML 2025 GenBio Workshop
[Paper]
[Code]
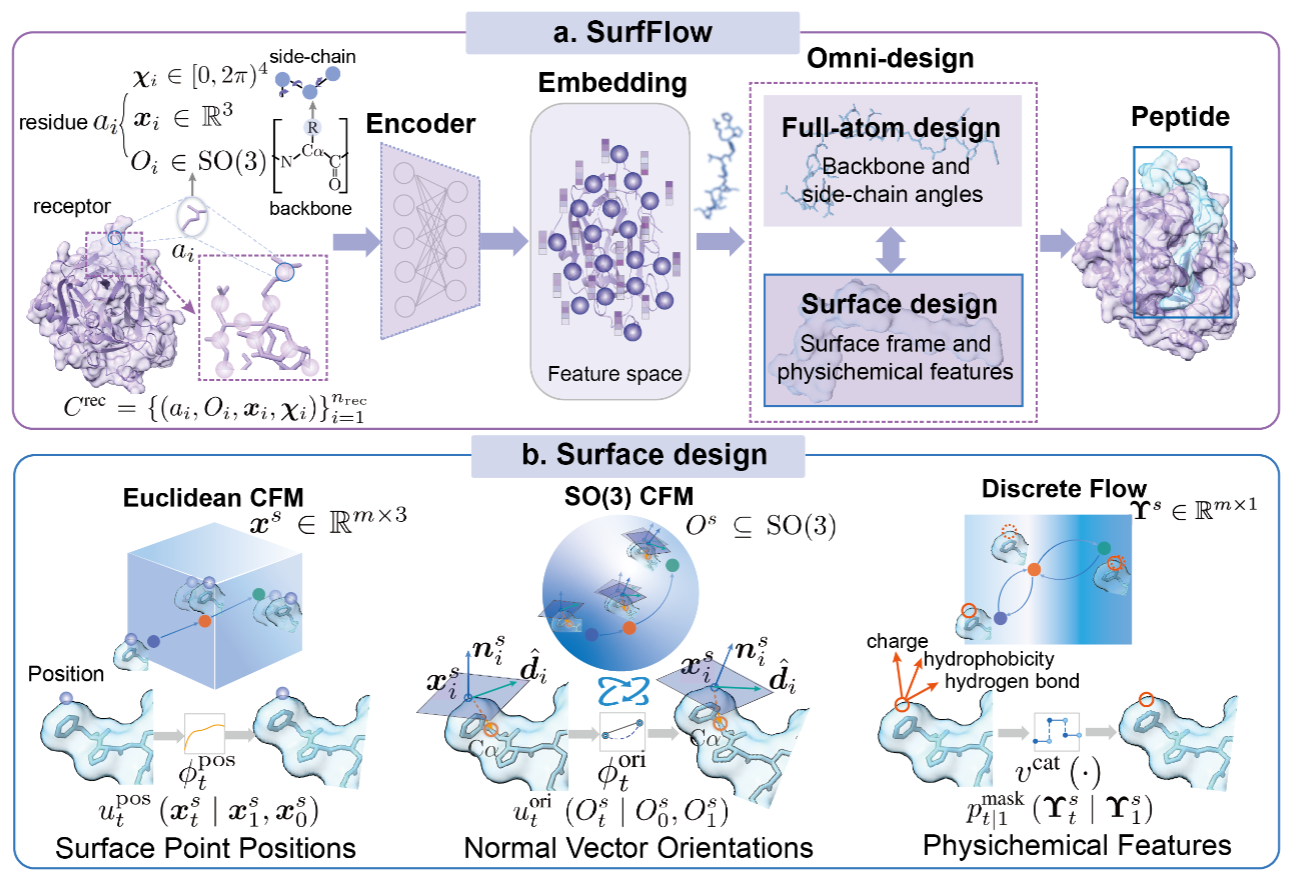 Surface-based Peptide Design with Multi-modal Flow Matching.
Surface-based Peptide Design with Multi-modal Flow Matching.
Fang Wu*, Shuting Jin, Zhengyuan Zhou, Xiangxiang Zeng, Jure Leskovec, Jinbo Xu†
KDD 2025
[Paper]
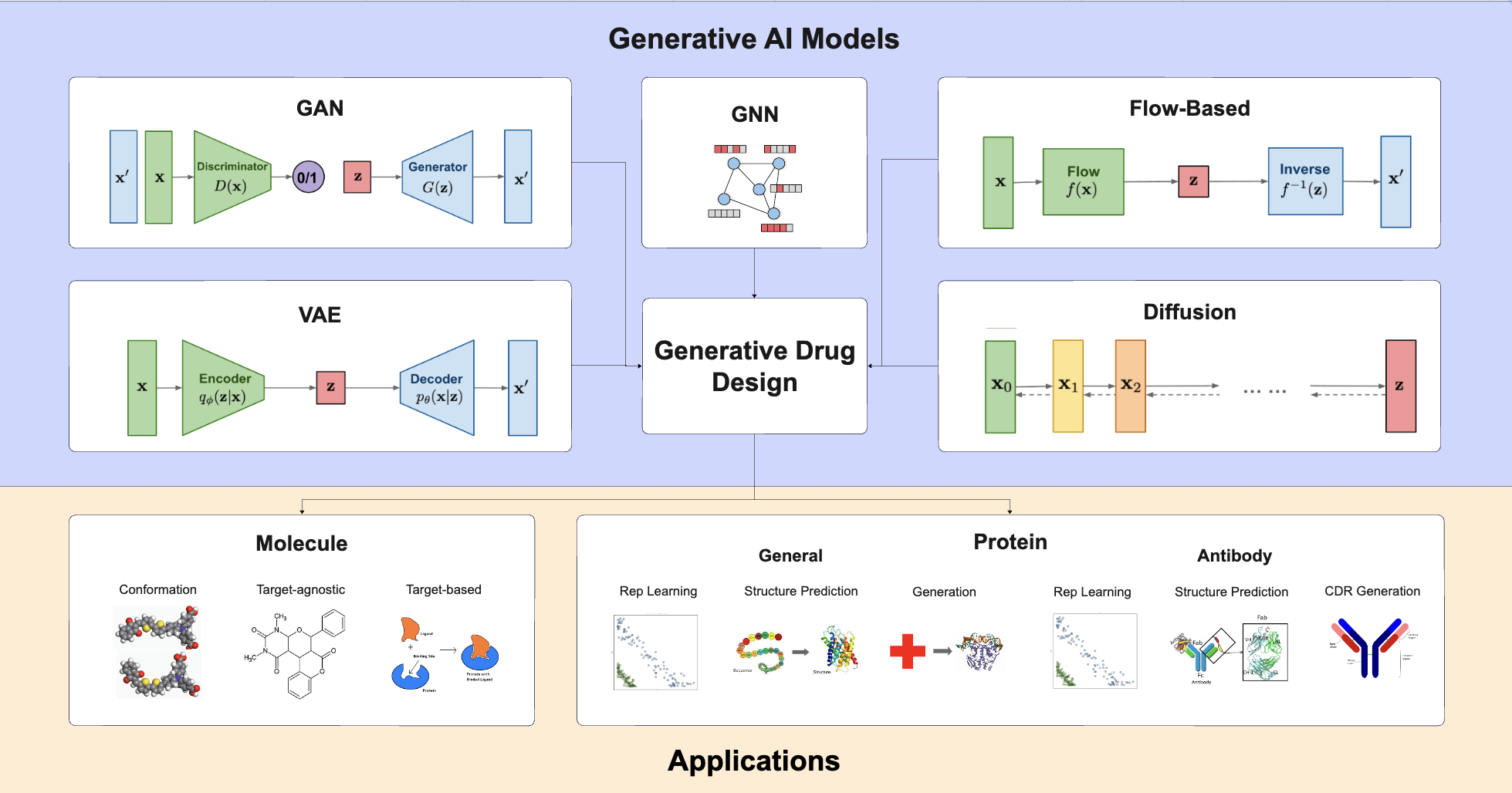 A Survey of Generative AI for de novo Drug Design: New Frontiers in Molecule and Protein Generation.
A Survey of Generative AI for de novo Drug Design: New Frontiers in Molecule and Protein Generation.

Xiangru Tang*, Howard Dai*, Elizabeth Knight*, Fang Wu,, Yunyang Li, Tianxiao Li, Mark Gerstein†
Briefings in Bioinformatics (2024)
[Paper]
[Github Repo.]
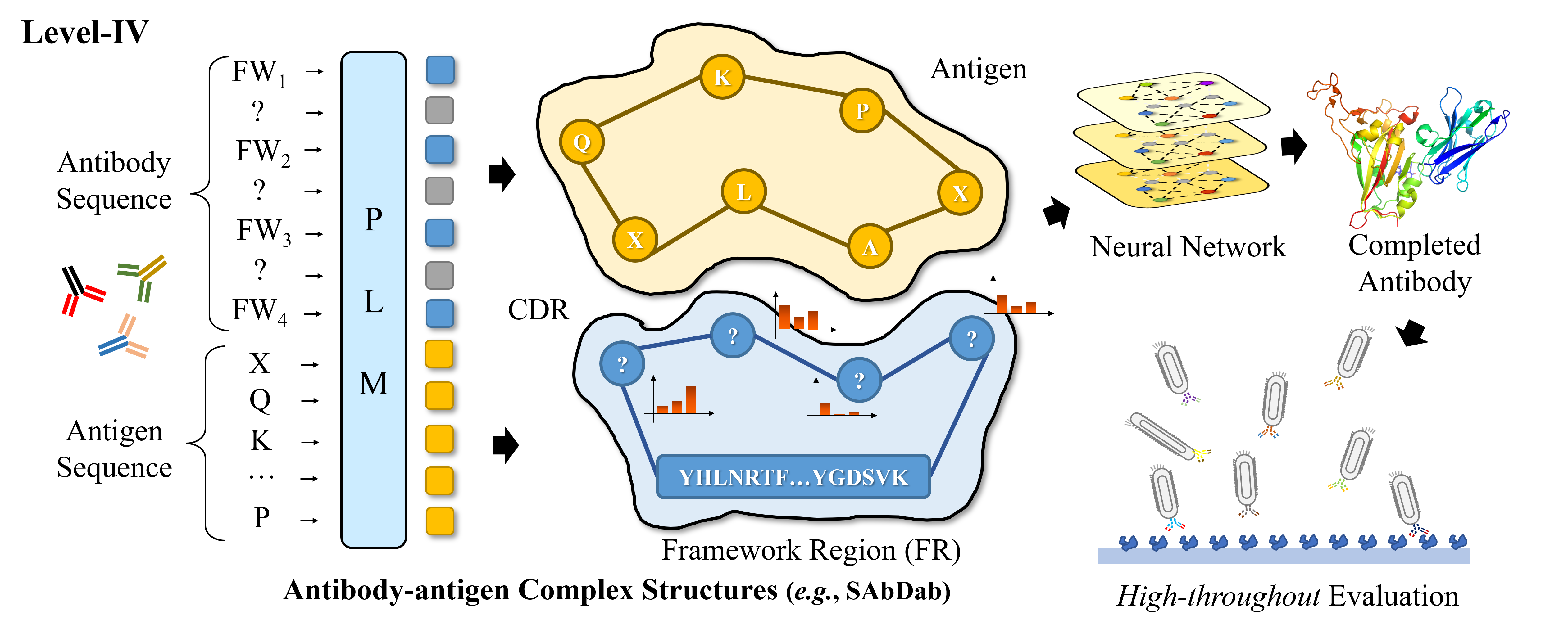 A Hierarchical Training Paradigm for Antibody Structure-sequence Co-design
A Hierarchical Training Paradigm for Antibody Structure-sequence Co-design
Fang Wu, Stan Z. Li†
NeurIPS 2023
[Paper]
 PoseX: AI Defeats Physics Approaches on Protein-Ligand Cross Docking.
PoseX: AI Defeats Physics Approaches on Protein-Ligand Cross Docking.

Yize Jiang*, Xinze Li*, Yuanyuan Zhang*, Jin Han*, Youjun Xu*, Ayush Pandit, Zaixi Zhang, Mengdi Wang, Mengyang Wang, Chong Liu, Guang Yang, Yejin Choi, WuJun Li†, Tianfan Fu†, Fang Wu†, Junhong Liu†
ICLR 2026
[Paper]
[Code]
[Webpage]
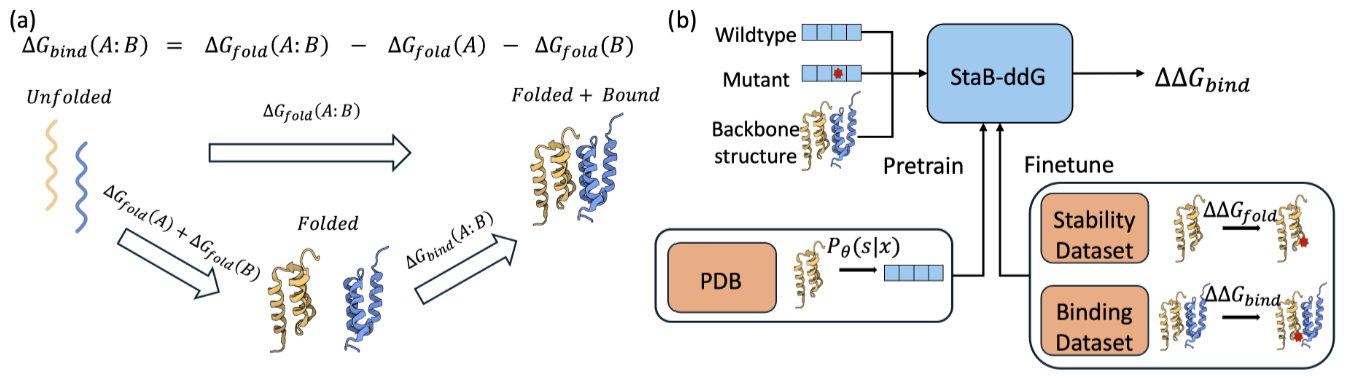 StaB-ddG: Predicting mutational effects on protein binding from folding energy.
StaB-ddG: Predicting mutational effects on protein binding from folding energy.

Arthur Deng, Karsten D. Householder, Fang Wu, Sebastian Thrun, K. Christopher Garcia, Brian L. Trippe†
ICML 2025
[Paper]
[Code]
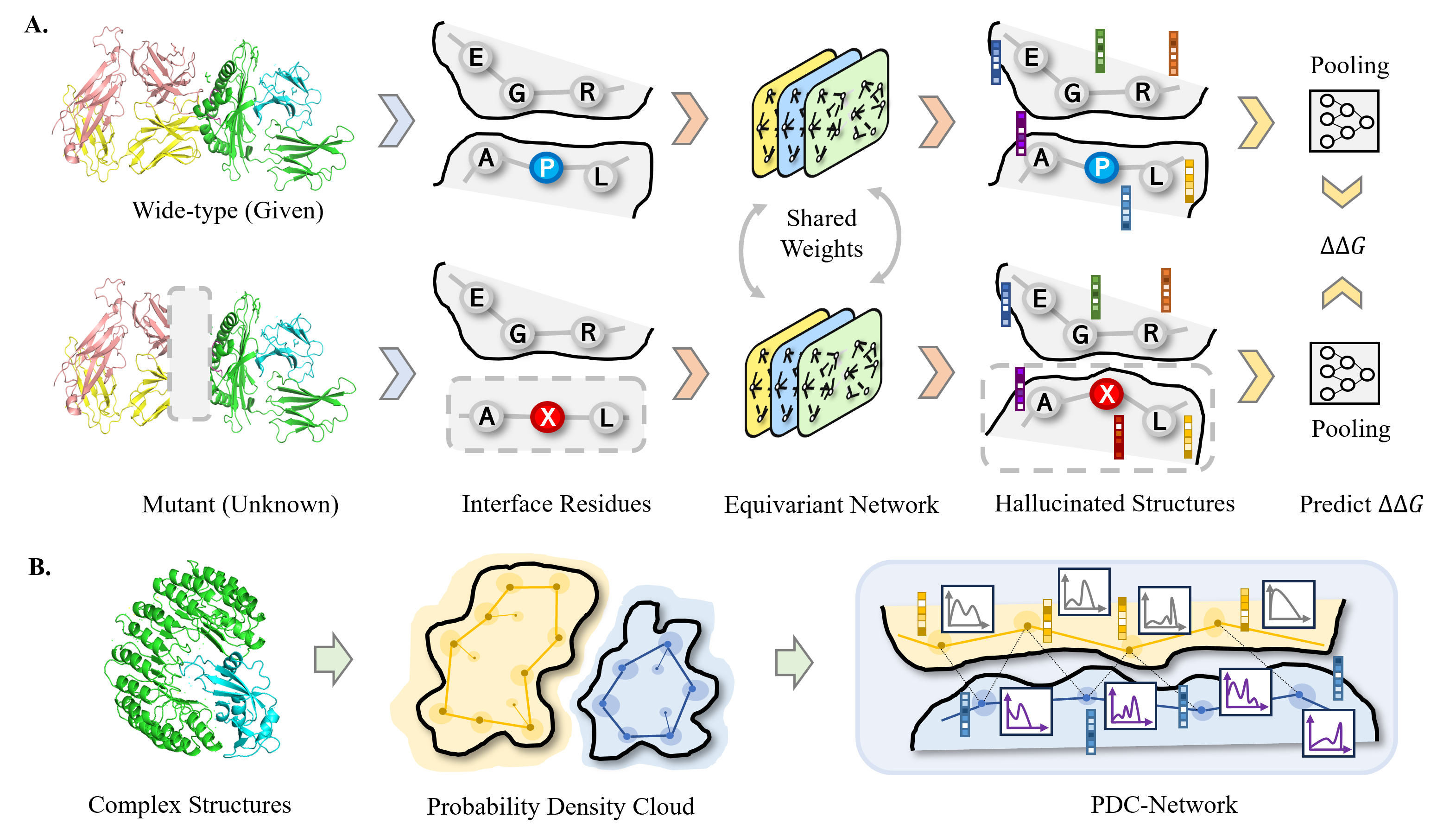 Dynamics-inspired Structure Hallucination for Protein-protein Interaction Modeling.
Dynamics-inspired Structure Hallucination for Protein-protein Interaction Modeling.

Fang Wu, Stan Z. Li†
TMLR (2025) → ICLR 2026 (Journal-to-Conference Track)
[Paper]
[Code]
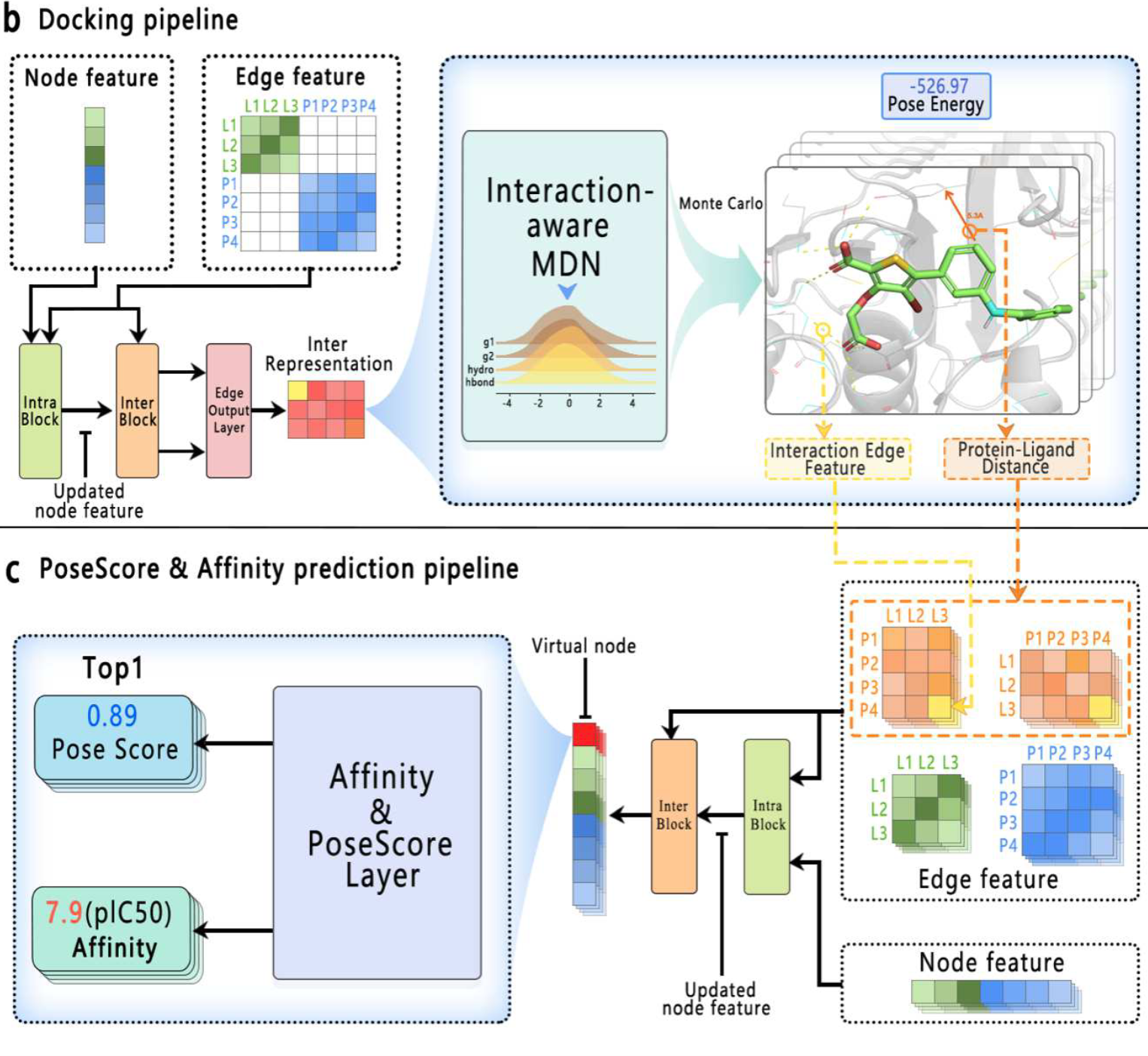 Interformer: An Interaction-Aware Model for Protein-Ligand Docking and Affinity Prediction.
Interformer: An Interaction-Aware Model for Protein-Ligand Docking and Affinity Prediction.

Houtim Lai†, Longyue Wang†, Ruiyuan Qian, Juhong Huang, Peng Zhou, Geyan Ye, Fandi Wu, Fang Wu, Xiangxiang Zeng, Wei Liu
Nature Communications (2024)
[Paper]
[Code]
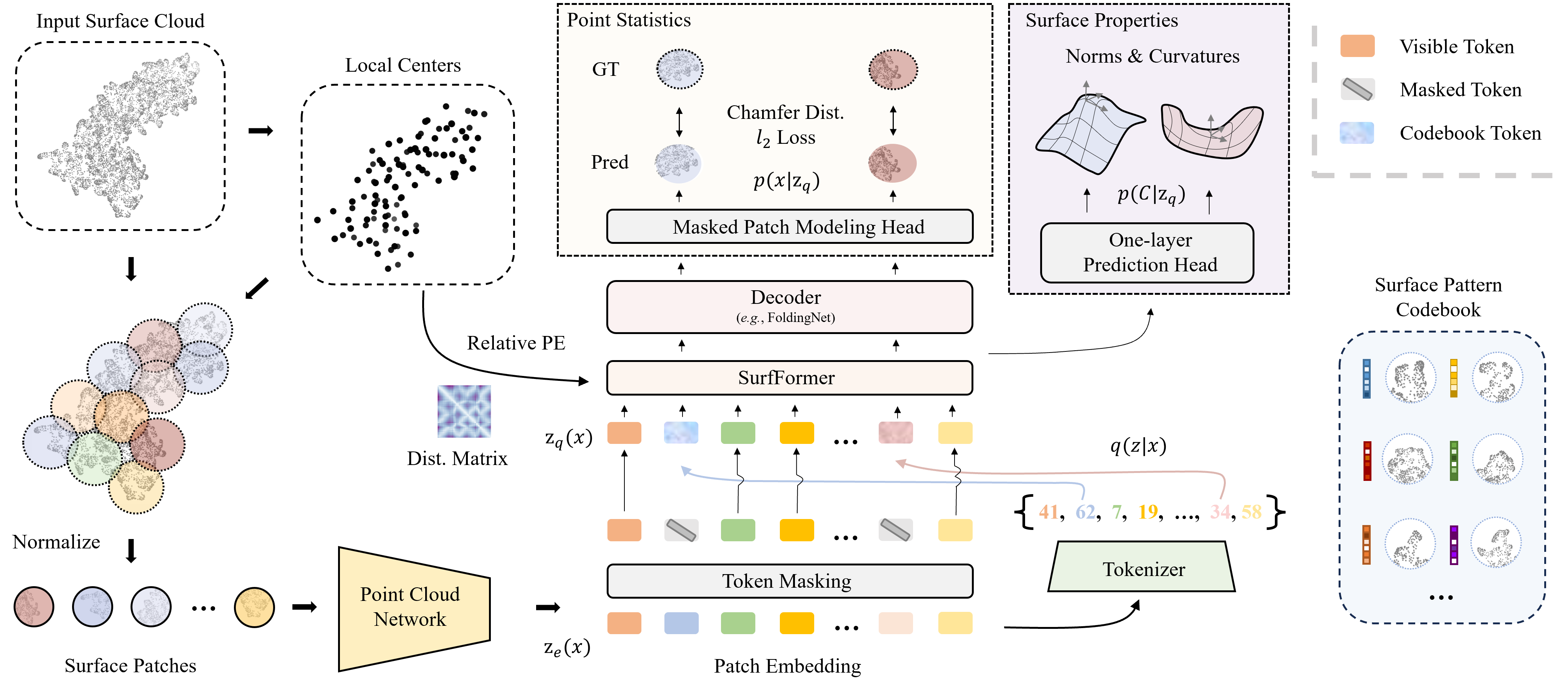 Surface-VQMAE: Vector-quantized Masked Auto-encoders on Molecular Surfaces.
Surface-VQMAE: Vector-quantized Masked Auto-encoders on Molecular Surfaces.

Fang Wu, Stan Z. Li†
ICML 2024
[Paper]
[Code]
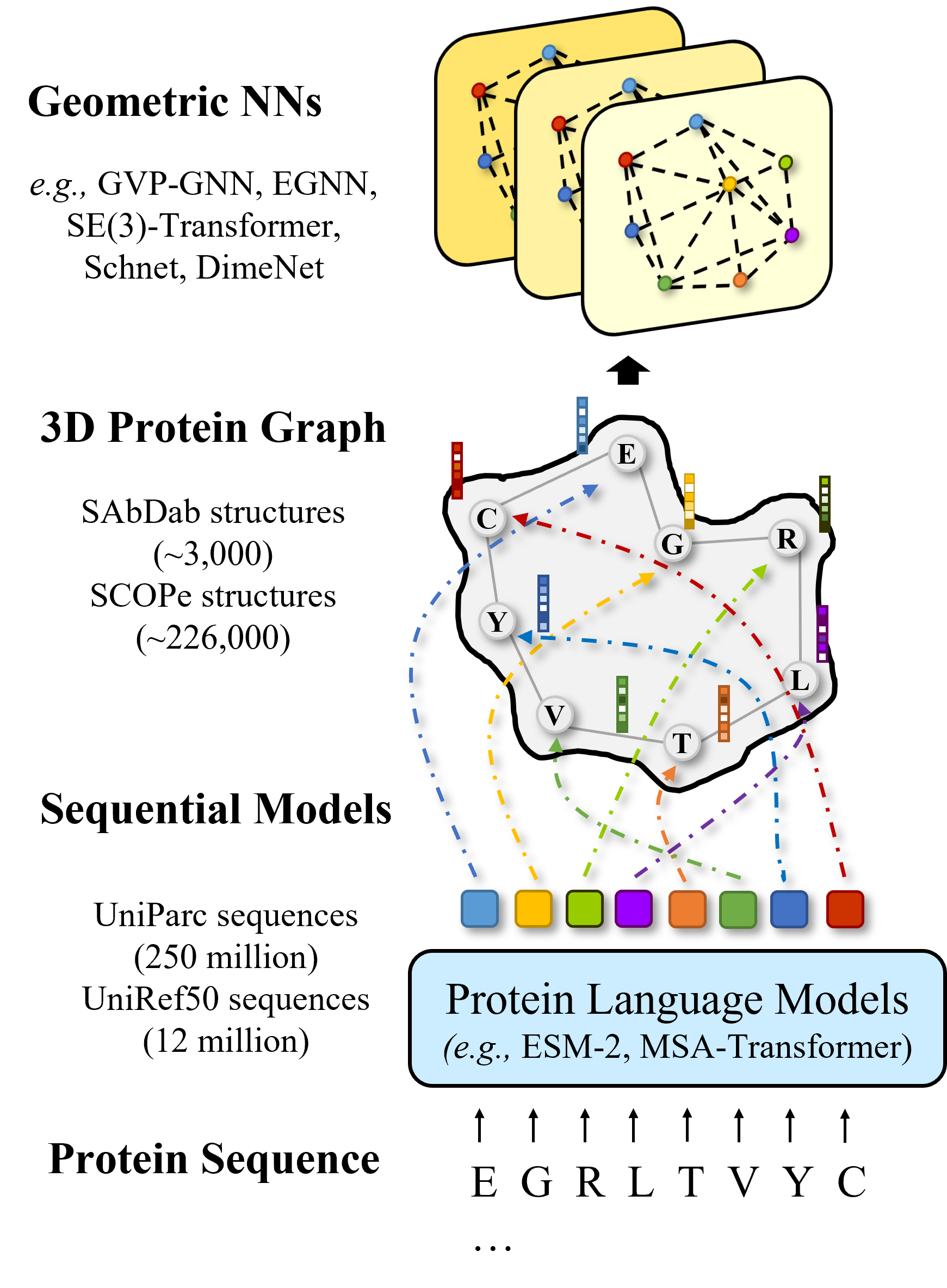 Integration of Pre-trained Protein Language Models into Geometric Deep Learning Networks.
Integration of Pre-trained Protein Language Models into Geometric Deep Learning Networks.

Fang Wu, Liong Wu, Dragomir Radev, Jinbo Xu, Stan Z. Li†
Communications Biology (2023)
[Paper]
[Code]
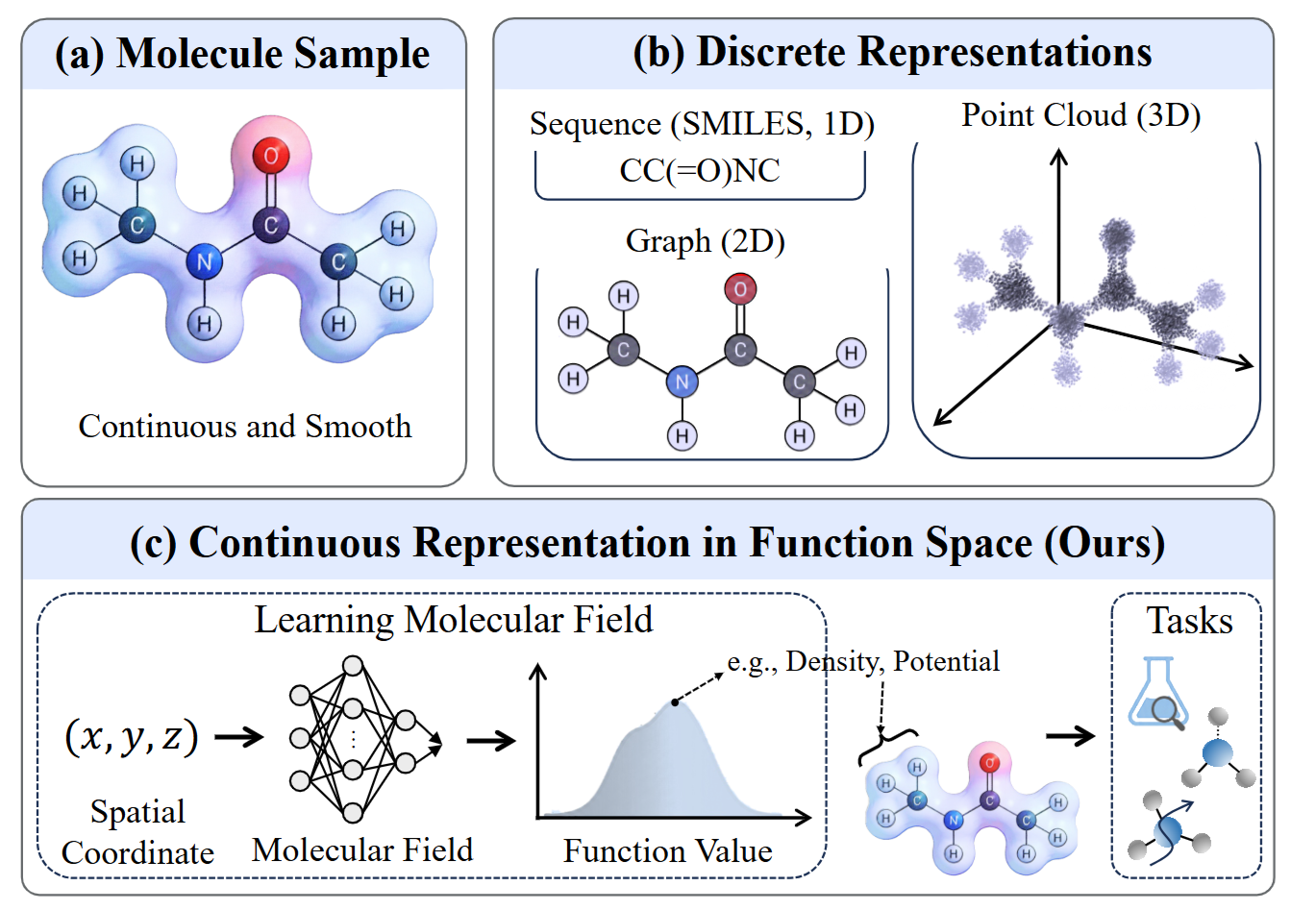 Molecular Representations in Implicit Functional Space via Hyper-Networks.
Molecular Representations in Implicit Functional Space via Hyper-Networks.

Zehong Wang, Xiaolong Han, Qi Yang, Xiangru Tang, Fang Wu, Xiaoguang Guo, Weixiang Sun, Tianyi Ma, Pietro Lio, Le Cong, Sheng Wang, Chuxu Zhang, Yanfang Ye†
Under Review
[Paper]
[Code]
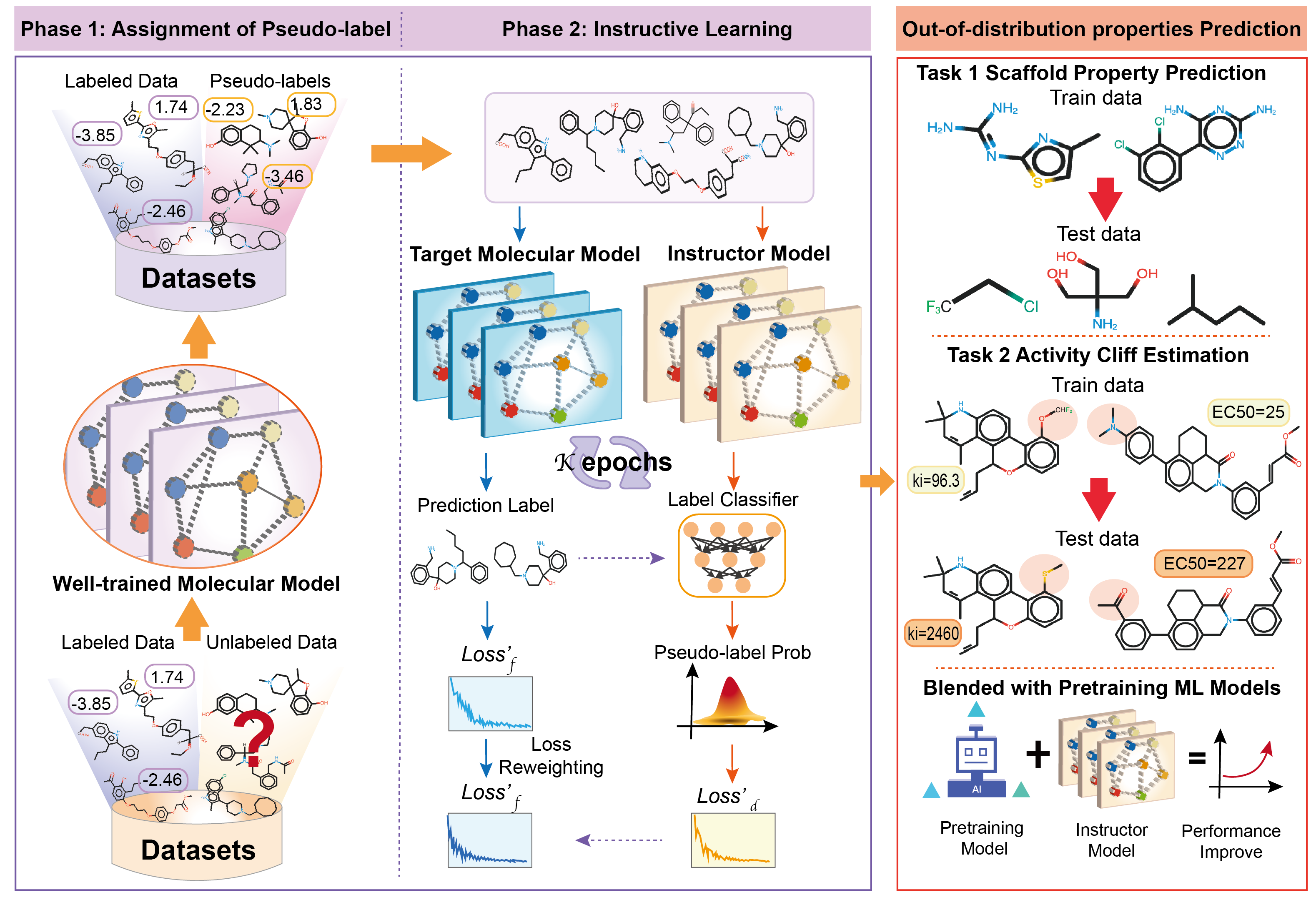 Instructor-inspired Machine Learning for Robust Molecular Property Prediction.
Instructor-inspired Machine Learning for Robust Molecular Property Prediction.

Fang Wu*†, Shuting Jin*, Siyuan Li, Stan Z. Li
NeurIPS 2024
[Paper]
[Code]
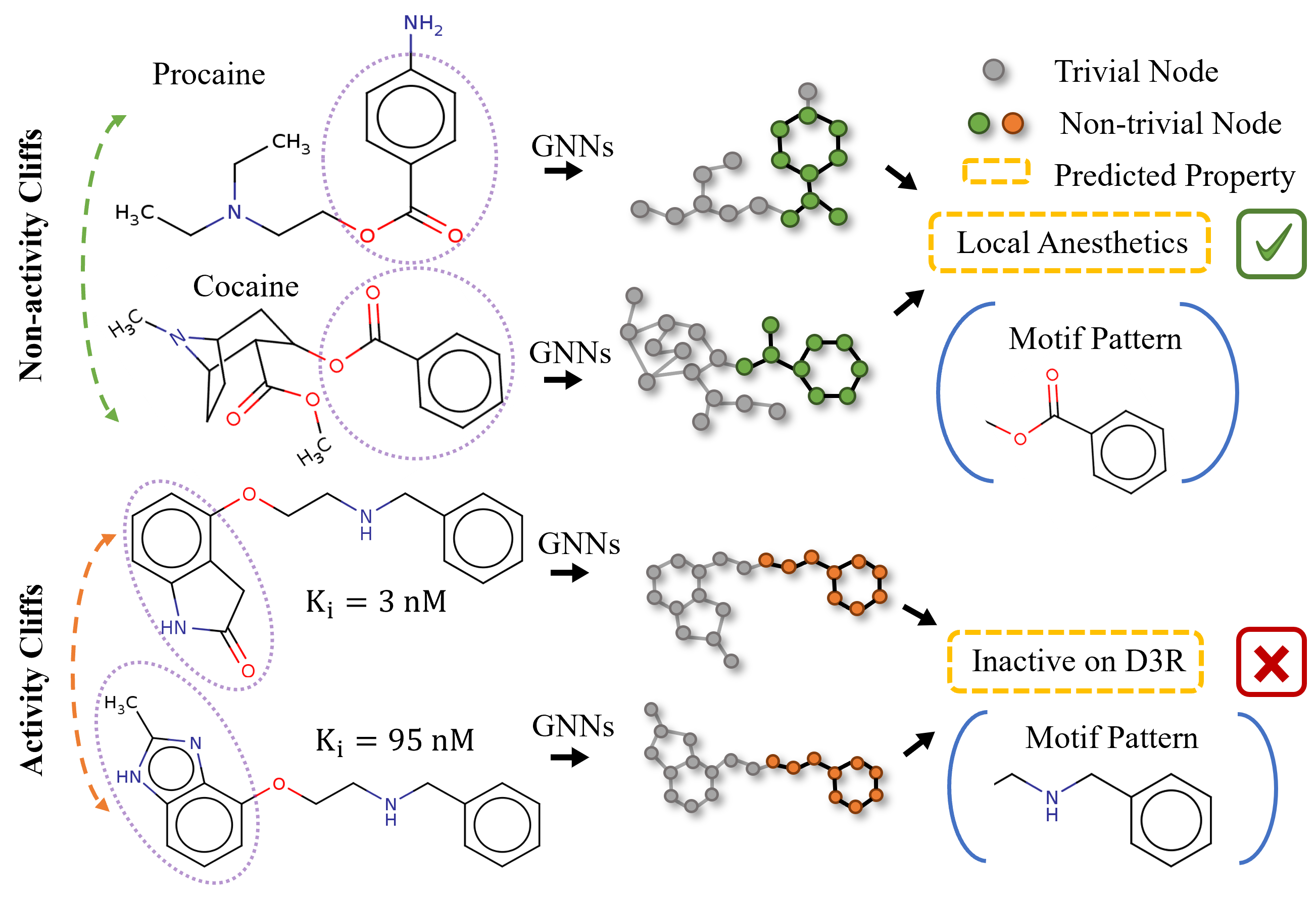 A Semi-supervised Molecular Learning Framework for Activity Cliff Estimation.
A Semi-supervised Molecular Learning Framework for Activity Cliff Estimation.

Fang Wu†
IJCAI 2024
[Paper]
[Code]
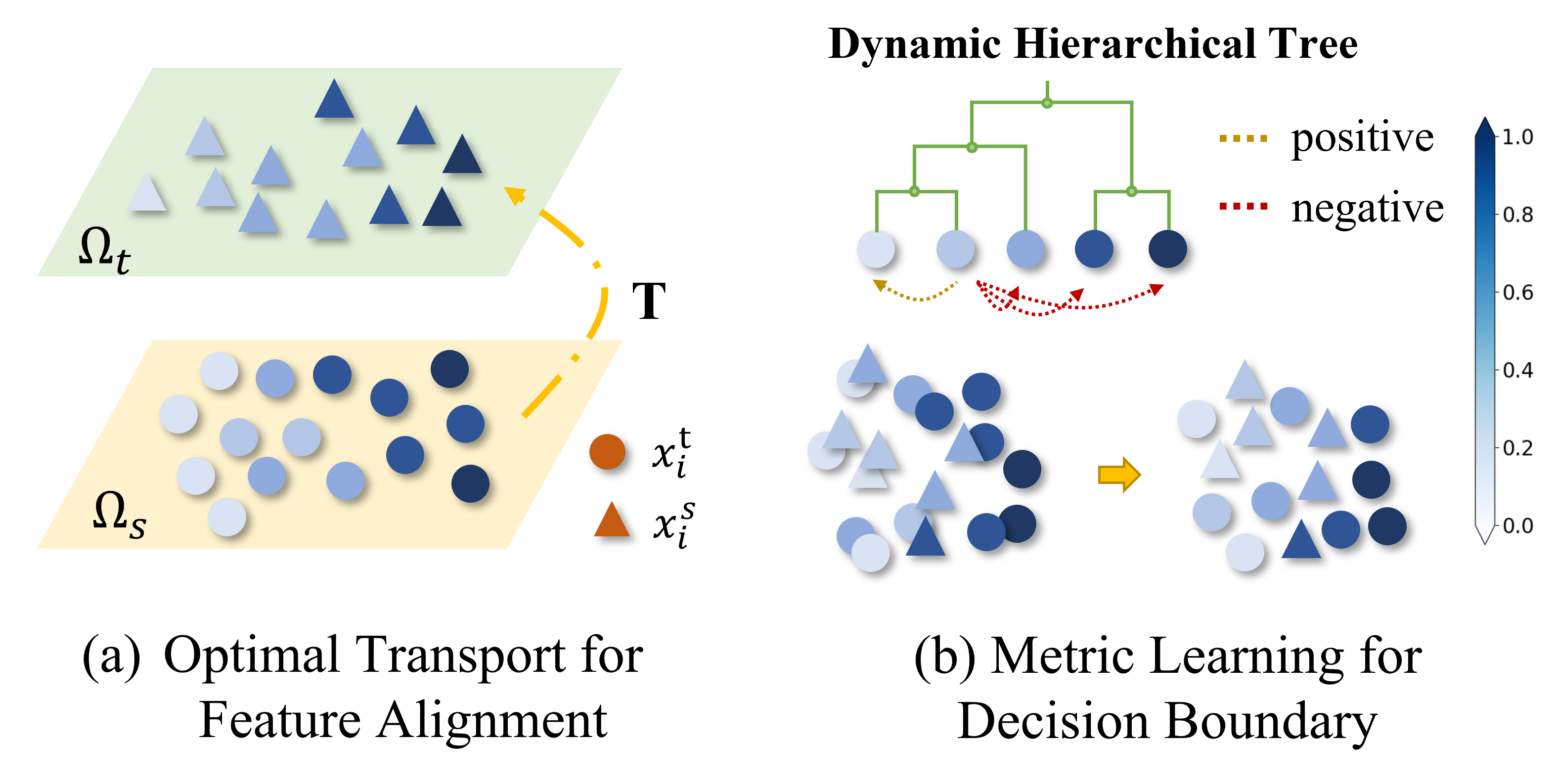 Improving Molecular Representation Learning with Metric Learning-enhanced Optimal Transport
Improving Molecular Representation Learning with Metric Learning-enhanced Optimal Transport

Fang Wu*, Nicolas Courty*, Shuting Jin*, Stan Z. Li†
Patterns (2023)
[Paper]
[Code]
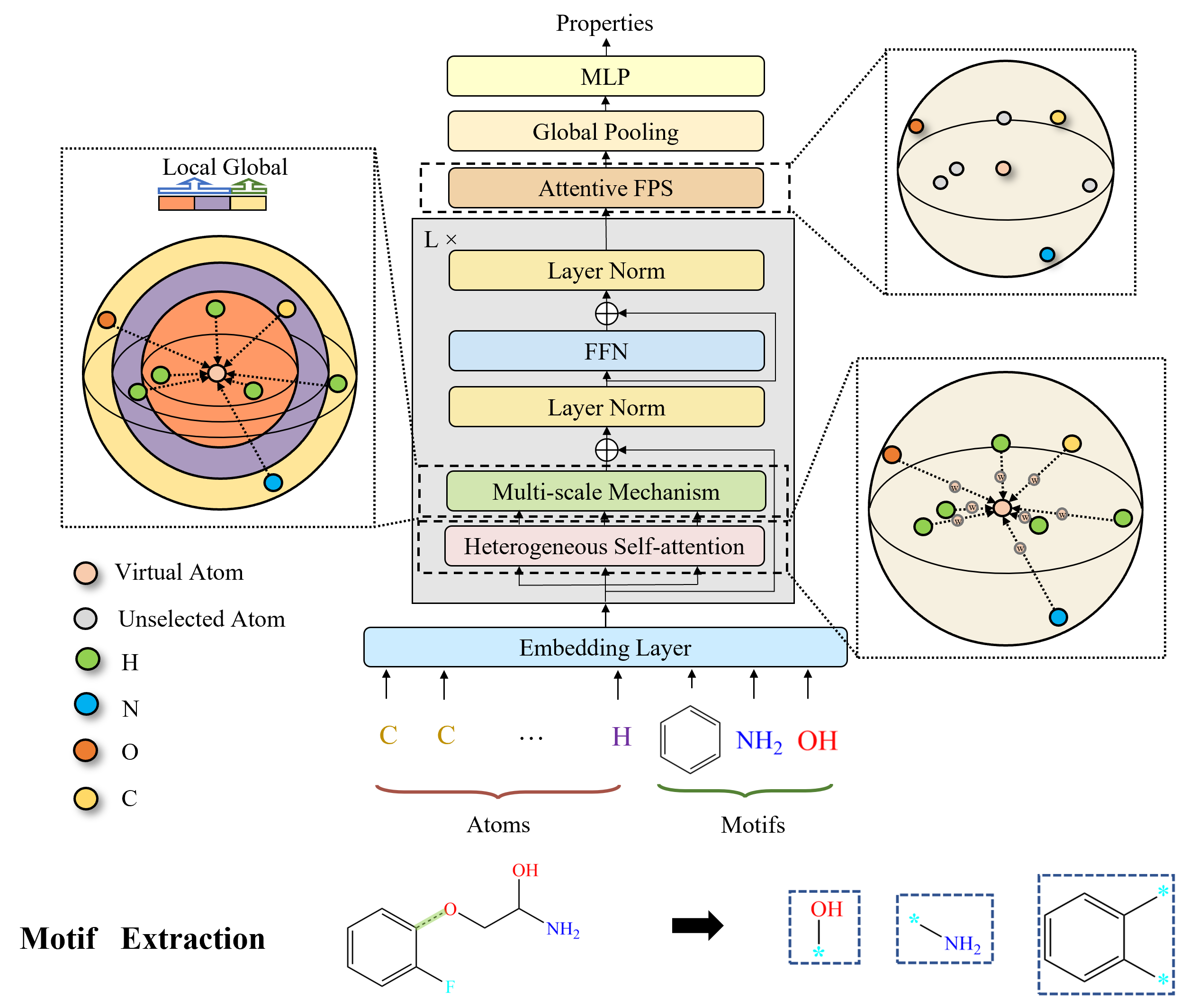 Molformer: Motif-based Transformer on 3D Heterogeneous Molecular Graphs.
Molformer: Motif-based Transformer on 3D Heterogeneous Molecular Graphs.

Fang Wu, Dragomir Radev, Stan Z. Li†
AAAI 2023 (oral)
[Paper]
[Code]
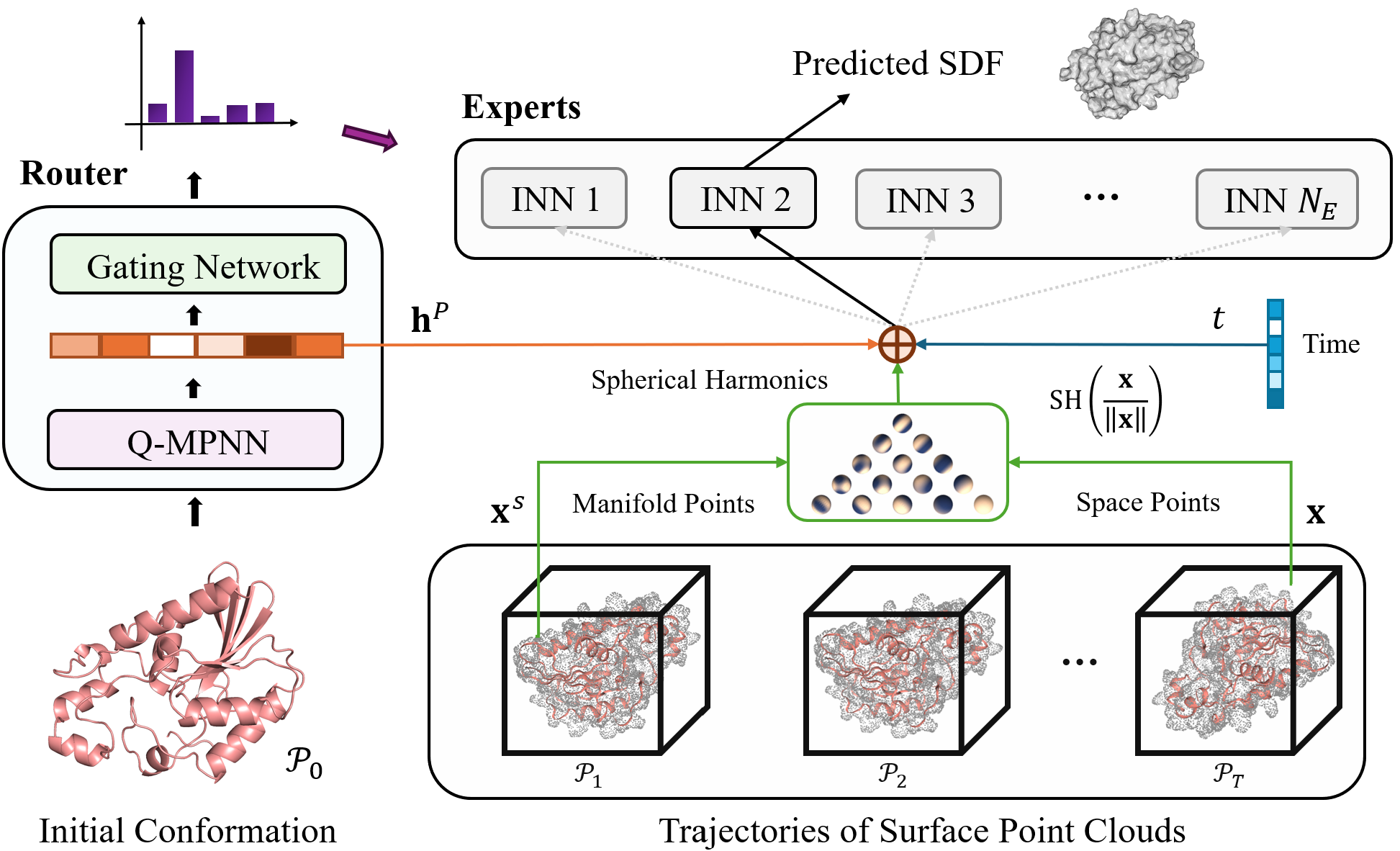 Generalized Implicit Neural Representations for Dynamic Molecular Surface Modeling.
Generalized Implicit Neural Representations for Dynamic Molecular Surface Modeling.
Fang Wu, Bozhen Hu, Stan Z. Li†
AAAI 2025
[Paper]
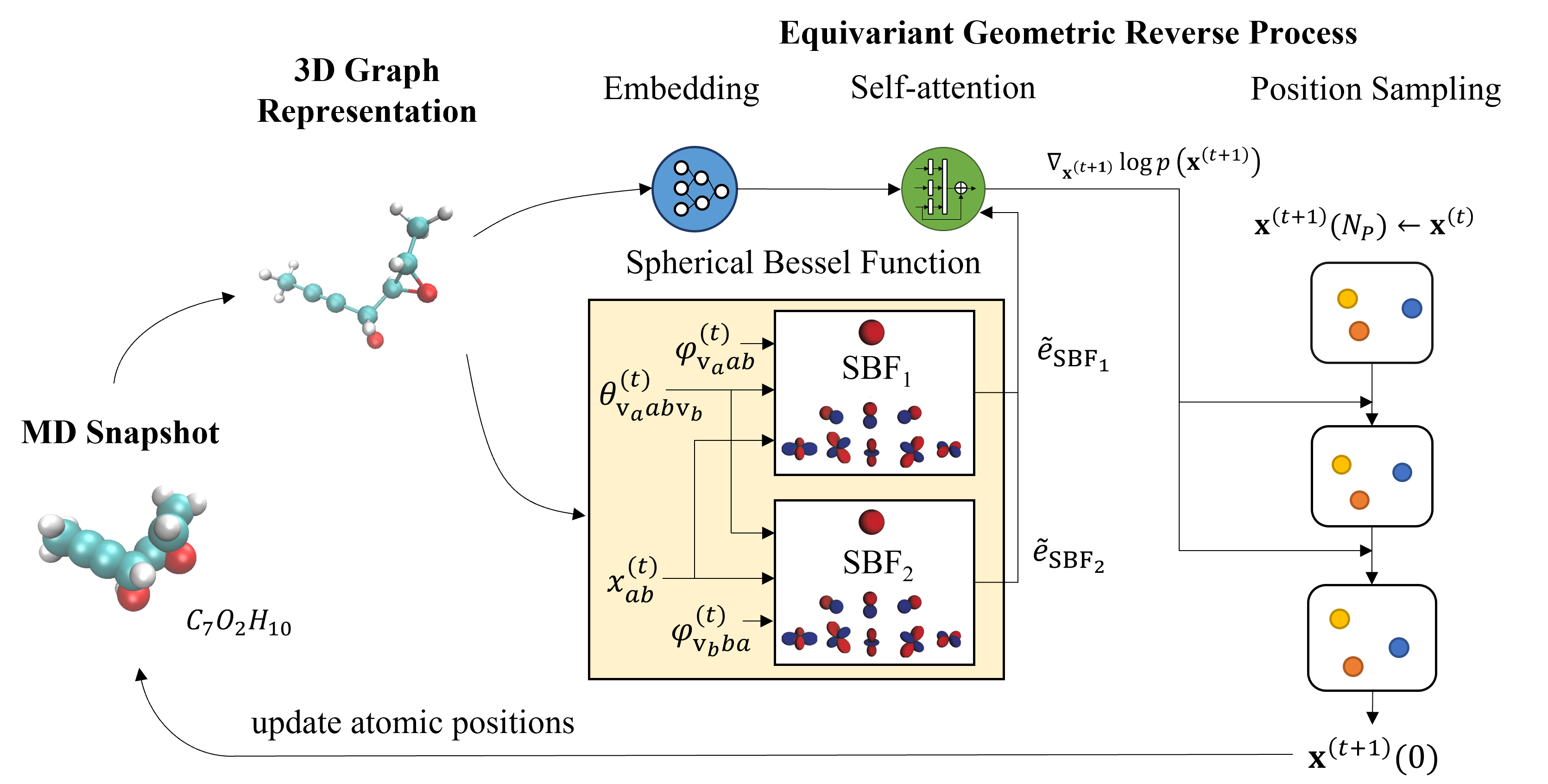 DiffMD: A Geometric Diffusion Model for Molecular Dynamics Simulations
DiffMD: A Geometric Diffusion Model for Molecular Dynamics Simulations
Fang Wu, Stan Z. Li†
AAAI 2023 (Oral)
[Paper]
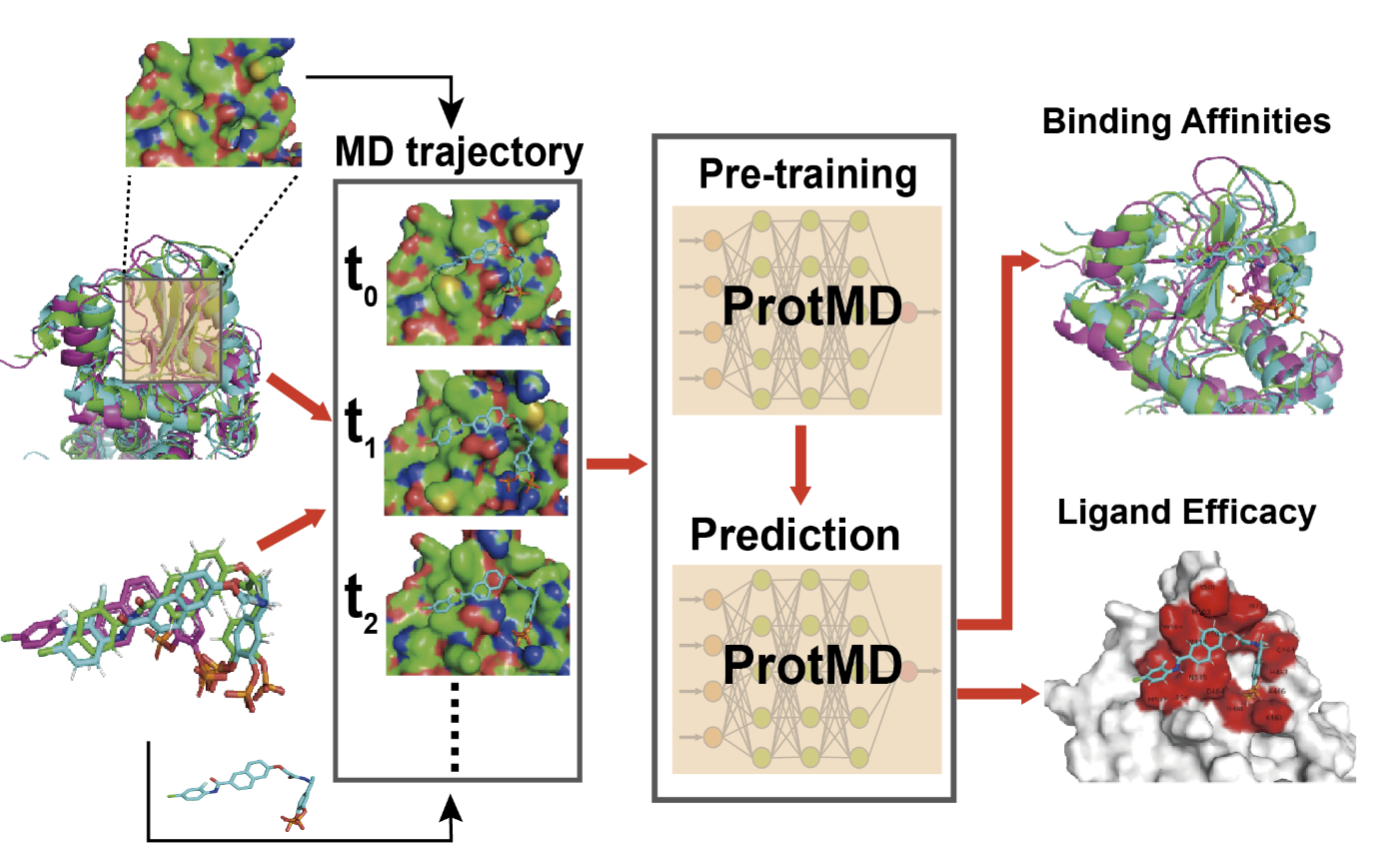 Pretraining of Equivariant Graph Matching Networks with Conformation Flexibility for Drug Binding
Pretraining of Equivariant Graph Matching Networks with Conformation Flexibility for Drug Binding

Fang Wu*, Shuting Jin*, Yinghui Jiang*, Xurui Jin, Bowen Tang, Zhangming Niu, Qiang Zhang, Xiangxiang Zeng, Stan Z. Li†
Advanced Science (2022)
[Paper]
[Code]
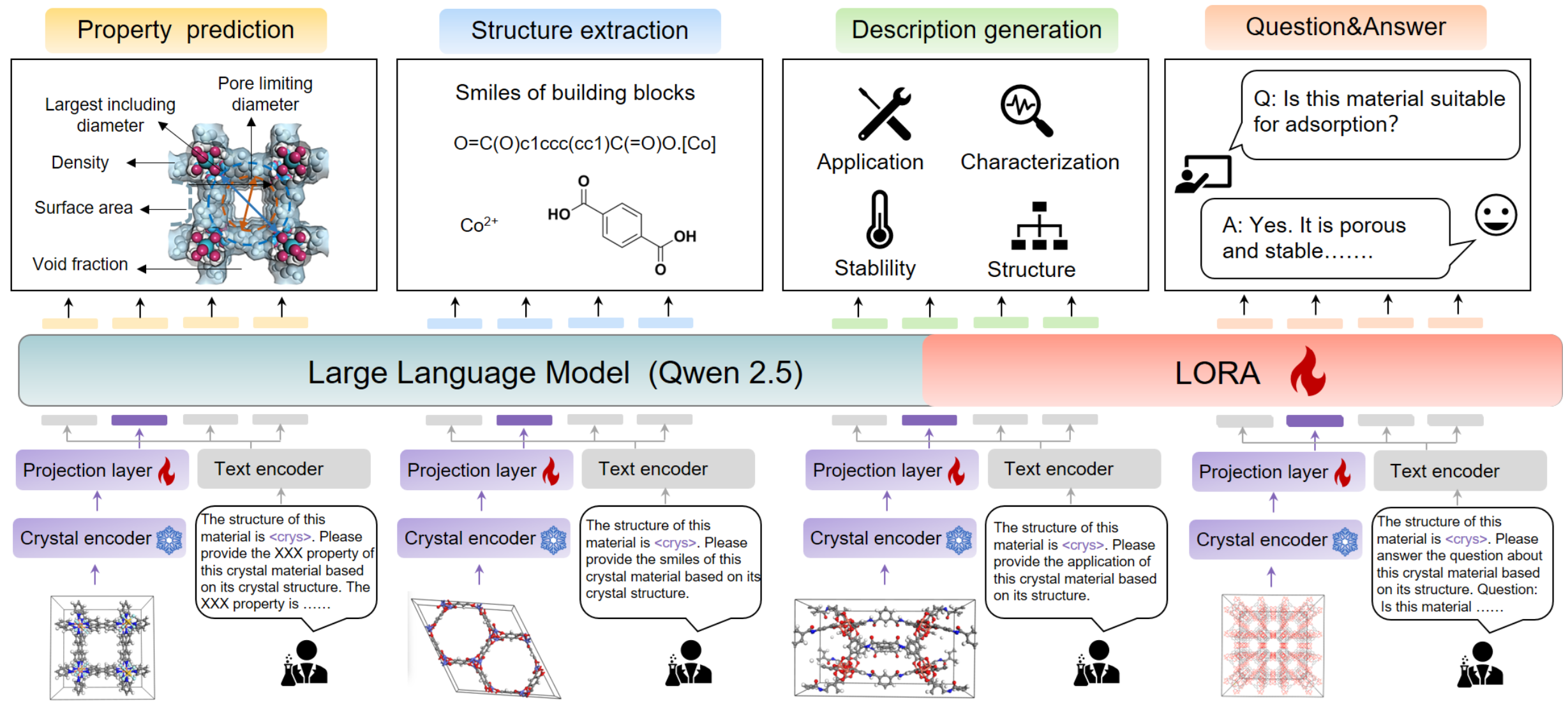 L2M3OF: A Large Language Multimodal Model for Metal-Organic Frameworks.
L2M3OF: A Large Language Multimodal Model for Metal-Organic Frameworks.

Jiyu Cui*, Fang Wu*, Haokai Zhao*, Minggao Feng, Xenophon Evangelopoulos, Andrew I. Cooper†, Yejin Choi†
Under Review
[Paper]
[Code]
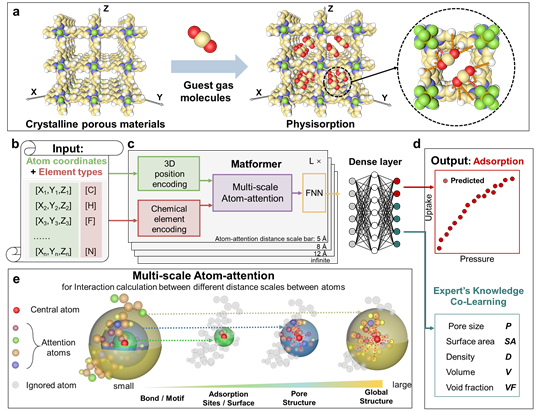 Direct Prediction of Gas Adsorption via Spatial Atom Interaction Learning.
Direct Prediction of Gas Adsorption via Spatial Atom Interaction Learning.

Jiyu Cui*, Fang Wu*, Wen Zhang*, Lifeng Yang*, Jianbo Hu, Yin Fang, Peng Ye, Qiang Zhang, Xian Suo, Yiming Mo, Xili Cui, Huajun Chen†, Huabin Xing†
Nature Communications (2023)
[Paper]
[Code]
 From Supervision to Exploration: What Does Protein Language Model Learn During Reinforcement Learning?
From Supervision to Exploration: What Does Protein Language Model Learn During Reinforcement Learning?  A Deep Reinforcement Learning Platform for Antibiotic Discovery.
A Deep Reinforcement Learning Platform for Antibiotic Discovery.
 Joint Design of Protein Surface and Structure Using a Diffusion Bridge Model.
Joint Design of Protein Surface and Structure Using a Diffusion Bridge Model.

 D-Flow: Multi-modality Flow Matching for D-peptide Design.
D-Flow: Multi-modality Flow Matching for D-peptide Design.

 SurfDesign: Effective Protein Design on Molecular Surfaces.
SurfDesign: Effective Protein Design on Molecular Surfaces.  BC-Design: A Biochemistry-Aware Framework for High-Precision Inverse Protein Folding.
BC-Design: A Biochemistry-Aware Framework for High-Precision Inverse Protein Folding.

 Surface-based Peptide Design with Multi-modal Flow Matching.
Surface-based Peptide Design with Multi-modal Flow Matching.  A Survey of Generative AI for de novo Drug Design: New Frontiers in Molecule and Protein Generation.
A Survey of Generative AI for de novo Drug Design: New Frontiers in Molecule and Protein Generation.

 A Hierarchical Training Paradigm for Antibody Structure-sequence Co-design
A Hierarchical Training Paradigm for Antibody Structure-sequence Co-design  PoseX: AI Defeats Physics Approaches on Protein-Ligand Cross Docking.
PoseX: AI Defeats Physics Approaches on Protein-Ligand Cross Docking.

 StaB-ddG: Predicting mutational effects on protein binding from folding energy.
StaB-ddG: Predicting mutational effects on protein binding from folding energy.

 Dynamics-inspired Structure Hallucination for Protein-protein Interaction Modeling.
Dynamics-inspired Structure Hallucination for Protein-protein Interaction Modeling.

 Interformer: An Interaction-Aware Model for Protein-Ligand Docking and Affinity Prediction.
Interformer: An Interaction-Aware Model for Protein-Ligand Docking and Affinity Prediction.

 Surface-VQMAE: Vector-quantized Masked Auto-encoders on Molecular Surfaces.
Surface-VQMAE: Vector-quantized Masked Auto-encoders on Molecular Surfaces.

 Integration of Pre-trained Protein Language Models into Geometric Deep Learning Networks.
Integration of Pre-trained Protein Language Models into Geometric Deep Learning Networks.

 Molecular Representations in Implicit Functional Space via Hyper-Networks.
Molecular Representations in Implicit Functional Space via Hyper-Networks.

 Instructor-inspired Machine Learning for Robust Molecular Property Prediction.
Instructor-inspired Machine Learning for Robust Molecular Property Prediction.

 A Semi-supervised Molecular Learning Framework for Activity Cliff Estimation.
A Semi-supervised Molecular Learning Framework for Activity Cliff Estimation.
 Improving Molecular Representation Learning with Metric Learning-enhanced Optimal Transport
Improving Molecular Representation Learning with Metric Learning-enhanced Optimal Transport

 Molformer: Motif-based Transformer on 3D Heterogeneous Molecular Graphs.
Molformer: Motif-based Transformer on 3D Heterogeneous Molecular Graphs.

 Generalized Implicit Neural Representations for Dynamic Molecular Surface Modeling.
Generalized Implicit Neural Representations for Dynamic Molecular Surface Modeling.  DiffMD: A Geometric Diffusion Model for Molecular Dynamics Simulations
DiffMD: A Geometric Diffusion Model for Molecular Dynamics Simulations  Pretraining of Equivariant Graph Matching Networks with Conformation Flexibility for Drug Binding
Pretraining of Equivariant Graph Matching Networks with Conformation Flexibility for Drug Binding

 L2M3OF: A Large Language Multimodal Model for Metal-Organic Frameworks.
L2M3OF: A Large Language Multimodal Model for Metal-Organic Frameworks.
 Direct Prediction of Gas Adsorption via Spatial Atom Interaction Learning.
Direct Prediction of Gas Adsorption via Spatial Atom Interaction Learning.

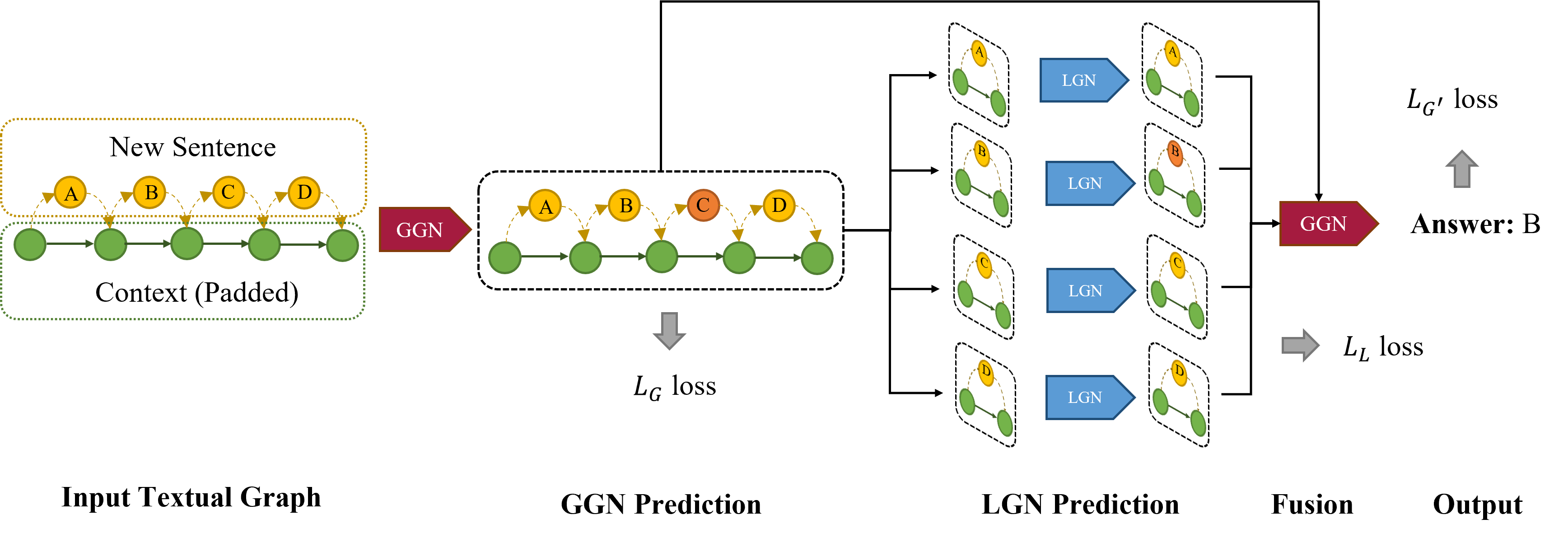 InsertGNN: A Hierarchical Graph Neural Network for the TOEFL Sentence Insertion Problem
InsertGNN: A Hierarchical Graph Neural Network for the TOEFL Sentence Insertion Problem

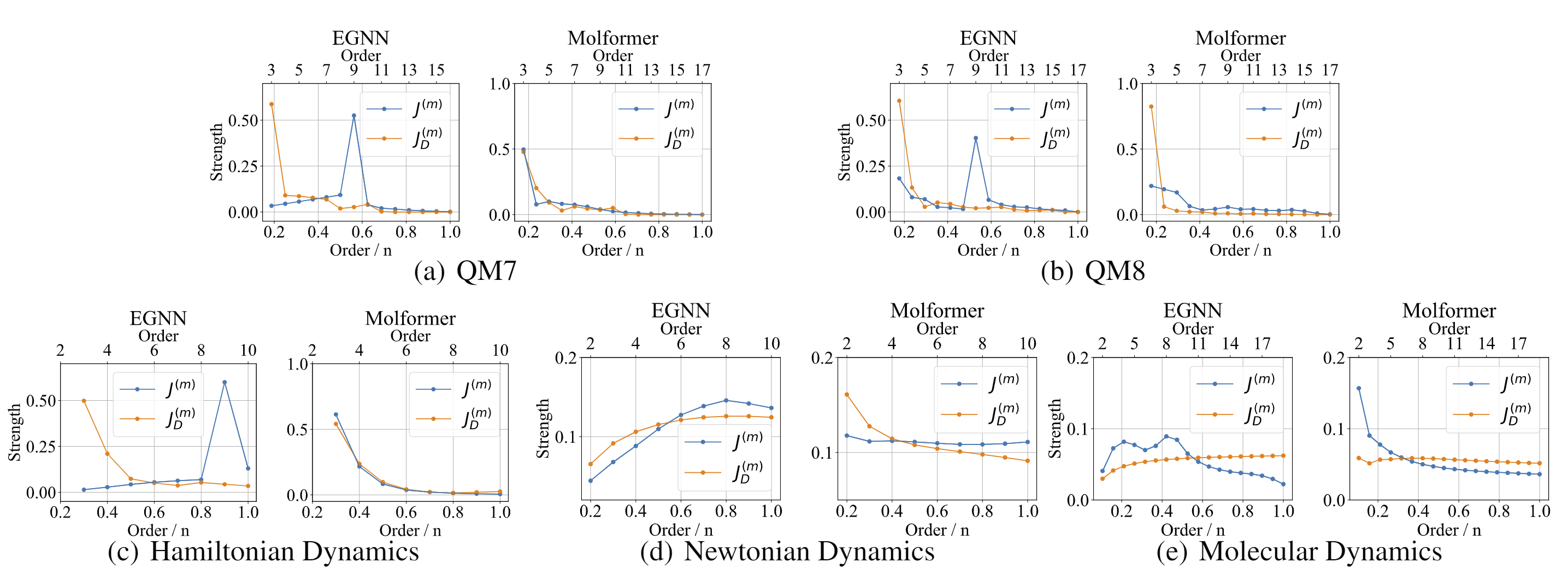 Discovering the Representation Bottleneck of Graph Neural Networks
Discovering the Representation Bottleneck of Graph Neural Networks

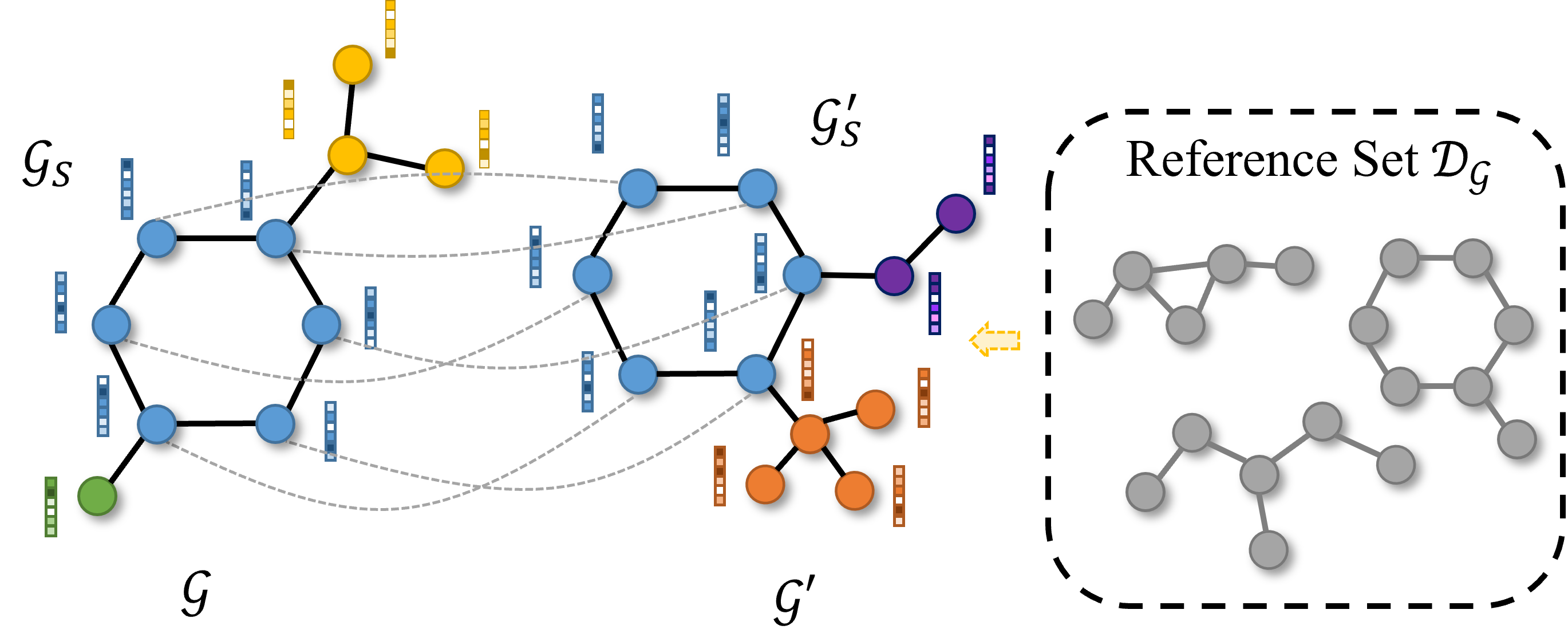 Rethinking Explaining Graph Neural Networks via Non-parametric Subgraph Matching
Rethinking Explaining Graph Neural Networks via Non-parametric Subgraph Matching
